| Issue |
Aquat. Living Resour.
Volume 34, 2021
|
|
|---|---|---|
| Article Number | 7 | |
| Number of page(s) | 10 | |
| DOI | https://doi.org/10.1051/alr/2021008 | |
| Published online | 02 April 2021 | |
Research Article
Can we identify wild-born salmon from parentage assignment data? A case study in the Garonne-Dordogne rivers salmon restoration programme in France
1
Université Paris-Saclay, INRAE, AgroParisTech, GABI, 78350 Jouy-en-Josas, France
2
MARBEC, Univ. Montpellier, CNRS, Ifremer, IRD, 34250 Palavas-les-Flots, France
3
SYSAAF Section Aquacole, INRAE LPGP, Campus de Beaulieu, 32042 Rennes, France
4
Labogena-DNA, Domaine de Vilvert, 78352 Jouy-en-Josas, France
5
Association Migado, 18ter rue de la Garonne, 47520 Le Passage d'Agen, France
* Corresponding author: This email address is being protected from spambots. You need JavaScript enabled to view it.
Handling Editor: Carlos Saavedra
Received:
25
November
2020
Accepted:
8
March
2021
Abstract
Parentage assignment with genomic markers provides an opportunity to monitor salmon restocking programs. Most of the time, it is used to study the fate of hatchery-born fish in those programs, as well as the genetic impacts of restocking. In such analyses, only fish that are assigned to their parents are considered. In the Garonne-Dordogne river basin in France, native salmon have disappeared, and supportive breeding is being used to try to reinstate a self-sustained population. It is therefore of primary importance to assess the numbers of wild-born returning salmon, which could appear as wrongly assigned or not assigned, depending on the power of the marker set and on the size of the mating plan. We used the genotypes at nine microsatellites of the 5800 hatchery broodstock which were used from 2008 to 2014, and of 884 upstream migrating fish collected from 2008 to 2016, to assess our ability to identify wild-born salmon. We simulated genotypes of hatchery fish and wild-born fish and assessed how they were identified by the parentage assignment software Accurassign. We showed that 98.7% of the fish assigned within the recorded mating plan could be considered hatchery fish, while 93.3% of the fish in other assignment categories (assigned out of the mating plan, assigned to several parent pairs, not assigned) could be considered wild-born. Using a Bayesian approach, we showed that 31.3% of the 457 upstream migrating fish sampled from 2014 to 2016 were wild-born. This approach is thus efficient to identify wild-born fish in a restoration program. It remains dependent on the quality of the recording of the mating plan, which we showed was rather good (<5% mistakes) in this program. To limit this potential dependence, an increase in the number of markers genotyped (17 instead of 9) is now being implemented.
Key words: Parentage assignment / restocking / fisheries
© M. Vandeputte et al., by EDP Sciences 2021
 This is an Open Access article distributed under the terms of the Creative Commons Attribution License (https://creativecommons.org/licenses/by/4.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
This is an Open Access article distributed under the terms of the Creative Commons Attribution License (https://creativecommons.org/licenses/by/4.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
1 Introduction
The ability to identify the parents of an individual fish using multilocus genotypes has been a game changer in the management of both fisheries and aquaculture stocks (Vandeputte and Haffray, 2014; Steele et al., 2019). In aquaculture breeding, it enabled the use of pedigree information without investment in numerous family tanks, strongly improving the precision of estimated breeding values and the possibility to control inbreeding. In fisheries management and stock enhancement programs, tracing an individual's origin back to its parents, combined with traceability on where and when the offspring of those parents was released, gives opportunities to assess the efficiency of releasing fish in the wild at various sites and life stages for supplementation (McGinnity et al., 2003; Aykanat et al., 2014; Steele et al., 2019). Provided the set of markers used has sufficient assignment power (sensu Vandeputte, 2012, i.e. taking into account the number of potential parents), all the offspring of genotyped broodstock fish can be considered genetically “tagged”, as their parents can be identified with a very low error rate (Beacham et al., 2019). The advantages of genetical tagging over physical tagging are 1) that genetically tagged fish are intrinsically tagged, while physical tagging requires a minimum size at tagging and thus at release, and 2) that it is easier to genotype the majority of the broodstock than to individually tag a large proportion of the fish released, thus reducing the necessary sampling and tagging efforts (Steele et al., 2019).
One of the key requirements to identify the parents of an individual is that the genotype of the parents for the markers genotyped in the offspring are available. When parental genotypes are missing, there are two types of consequences. First, the immediate effect of missing parental genotypes is that the parents of the tested individual cannot be readily identified. Second, if either only one of the true parents, or some relatives of the parents, are present in the set of genotyped parents, it is likely that, in a significant proportion of cases, there may be a wrong identification of parents (false assignment) due to similarities between the genotype of the unknown true parent(s) and the genotype of the available parents (Griot et al., 2020). This is especially true if the assignment power of the marker set used is not very high. If assignment power is not high enough, it is also likely that the true parent pair may not be discriminated from other parents with relatively similar genotypes, leading to poly-assigned (potentially assigned to several parent pairs) offspring, which in the end has the same result: the true parents cannot be assigned with reasonable certainty.
The different cases are summarized in Table 1.
In most applications, the main aim is to maximize the rate of true positives while controlling the amount of false positives, so that the animals declared as “assigned” by the software are as reliably assigned as possible. Assignment software often gives the possibility to control Type I error a priori like CERVUS (Kalinowski et al., 2007) or APIS (Griot et al., 2020), which set a reliability threshold for assignments, or a posteriori like COLONY which associates a probability to each parent pair (Wang, 2012). Controlling Type II error is essentially necessary for cost reasons, because a high type II error implies a higher genotyping effort to achieve the same number of usable records. In general, in all applications, there is little interest for true negatives, and the way to avoid them is to ensure collecting DNA samples and genotyping of all potential parents. In the context of salmon restoration programs, hatchery juveniles released in the river are generally adipose fin clipped or tagged with a coded wire tag, and unclipped/untagged individuals are considered wild-born (Hess et al., 2012; Evans et al., 2015). Alternatively, in a few programs, all individuals are trapped at dams and sampled for DNA before being allowed to move to the spawning grounds, so that all parents of the wild-born individuals are also known (Araki et al., 2009), and thus wild-born individuals can be assigned as true positives. However, this obviously requires a large investment and depends on site equipment and morphology, and then cannot be applied in all programs. Moreover, some precocious male parr may mature and contribute to reproduction, escaping sampling if sampling is, as usually done, focused on migrating fish, thus further limiting the completeness of this approach (Aykanat et al., 2014).
The number of true negatives can be a key issue in the case of the genetic monitoring of a restoration programme where the wild population to be restored has been heavily depleted, or has even disappeared. In such programs, the final aim is to re-establish a self-sustaining population, and it is thus of primary importance to assess the proportion of fish that derive from natural reproduction, and hence from parents which are not hatchery broodstock. Indeed, there may be, in many cases, a positive relationship between fitness and population size, known as the Allee effect, which implies that a minimum population size is necessary for a population to be self-sustainable (Stephens and Sutherland, 1999; Kuparinen et al., 2014).
In France, Atlantic Salmon disappeared from the Garonne-Dordogne basin during the late 19th − early 20th century, due to the building of hydropower dams (Thibault, 1994). Following the establishment of fish passes, the first attempts to reintroduce Atlantic salmon in this river system date back to the 1980's, first with fish from Canada, Scotland and Norway, then in a second phase with fish from French origin (Loire-Allier and Adour), which resulted in the return of limited numbers of potentially spawning adults. Since 1995, a captive broodstock has been established by Association Migado, which manages the restoration programme. Each year, migrating adults (F0) are captured in the Garonne and Dordogne rivers, kept in a breeding center in Bergerac, then stripped to produce F1 offspring by artificial fertilization. The F1 fish are (1) released at different points of the two basins for direct restocking and (2) sent to multiplication hatcheries where they are grown to the broodstock stage to produce F2 offspring, which are then released in the wild at different stages (5% as eyed eggs, 90% as first-feeding fry, 3% as smolts and 2% as 1+ parr). Since 2008, all F0 migrants kept in Bergerac, and all F1 broodstock in the multiplication hatcheries have been genotyped for nine microsatellite markers. In addition, all crosses performed to produce the F1 and F2 families have been recorded. As hatchery fish are often released at very young stages (eyed eggs or first feeding fry), they cannot be tagged by adipose fin clipping or Coded Wire Tag. In addition, only a limited proportion of fish are sampled in the fish passes. Thus, the genotypes of potential wild parents are unknown.
We investigated the possibility to use parentage assignment data to qualify “wild-born” individuals, when only hatchery parents are genotyped, and hatchery offspring are not tagged. To this end, using real parental genotypes, we simulated the genotypes of F1 and F2 offspring from hatchery or non-hatchery parents, and examined how they were discriminated by the parentage assignment software Accurassign (Boichard et al., 2014) used to monitor the Garonne-Dordogne Atlantic salmon program. Using a Bayesian approach, we used these results to estimate the proportion of wild-born individuals among the 2014–2016 upstream migrating adults, and to assess the reliability of assigning a “hatchery” or “wild-born” origin to an individual, conditional on its qualification by the parentage assignment software.
Types of assignments and errors in parentage assignment with genomic markers, depending on the availability of parental genotypes.
2 Material and methods
2.1 Base data
The base data were the genotypes at nine microsatellites of a total of 5800 F0 and F1 hatchery broodstock used from 2008 to 2014, and of 884 upstream migrating fish collected from 2008 to 2016. The nine microsatellite markers used were SSOSL85 and SSOSL311 (Slettan et al., 1995), SSspG7, SSsp1605, SSsp2201, SSsp2210, SSsp2213, SSsp2215 and SSsp2216 (Paterson et al., 2004). Basic statistics and exclusion power of these markers are given in Table 2. Using combined Q3 and Q1 exclusion probabilities from Table 2, we inferred, following formula (7) in Vandeputte (2012), that the exclusion power of the marker set was 0.902 in a design with 3500 potential female parents and 2000 potential male parents, which is representative of what has to be resolved in this restoration programme (see below). The mating plans were recorded for all F0 and F1 hatchery crosses performed from 2008 to 2014.
Variability and assignment power of the nine microsatellite markers estimated from the F0 salmon broodstock present in Bergerac hatchery from 2010 to 2014 (156 individuals).
2.2 Simulation process
The aim of the simulation process was to generate genotypes which are representative of the salmon run of a given year, with a similar age structure, in order to assess how parents can be identified by the parentage assignment software.
In a given salmon run, there is a mixture of one sea-winter (1SW), two sea-winter (2SW) and three sea-winter (3SW) individuals. Reproduction happens in winter (December year N-1 to January year N), juveniles (parr) stay in the river and then migrate to the sea as smolts, generally in the spring of year N+1, but up to year N+3 for a small proportion of them. Migration back to the river happens in summer of year N+2 for 1SW salmon, and spring of years N+3 and N+4 for 2SW and 3SW salmon, respectively. Thus, in the 2014 salmon run, 1SW salmon are mostly from the 2012 winter reproduction season, 2SW from 2011 and 3SW from 2010. A small proportion of animals, having spent 2 or 3 year in the river, might be from the 2008 and 2009 reproduction seasons. Thus, parentage of individuals from the 2014 salmon run thus has to be tested on all hatchery broodstock used in the 2008–2012 reproduction seasons.
For every salmon run, there is a specific proportion of 1SW, 2SW and 3SW fish. The proportions for the 2014–2016 runs are given in Appendix A.
For each of those three run years, we simulated potential offspring genotypes from four different origins:
-
F1 fish from F0 parents, from Bergerac hatchery
-
F2 fish from F1 parents born in Bergerac, from Castels hatchery
-
F2 fish from F1 parents born in Bergerac, from Pont-Crouzet-Cauterêts hatchery
-
Wild-born individuals from F0 parents sampled in fish traps but not collected to renew the F0 stock of Bergerac.
We considered that, as the vast majority of young salmon migrate to the sea at 1 year, only the mating plans of years N-2, N-3 and N-4 would be used to generate offspring. In all hatcheries, the typical mating plan is a series of small factorial designs, each performed on a given day. Statistics on the mating plans are given in Table 3. In a given year, in general females are used in two factorial designs in Bergerac, and in one factorial design in F1 hatcheries, while males are used on average in 30 factorials in Bergerac, and in only one factorial in F1 hatcheries.
For each salmon run, 1000 individuals were simulated from each hatchery, using an in-house VBA script in Microsoft Excel (provided as Supplementary Material 1). For each individual from that hatchery, the simulation process was the following: (1) a year of birth was assigned to the individual following the distributions of 1SW, 2SW and 3SW fish corresponding to that salmon run year (Appendix A) (2) a factorial cross was randomly chosen among the ones performed that year in that hatchery, (3) a male and a female were randomly chosen among the ones in that factorial and (4) for each locus, one allele from the male and one allele from the female were randomly chosen to obtain the offspring's genotype. The real mating plans described in Table 3 were used as the basis for these simulations.
For wild-born individuals, the process was the same, with 1000 offspring generated, except that the “broodstock” of year N was composed of wild individuals sampled at fish traps in year N-1, that were genotyped but released to the river after sampling and thus not used to renew the Bergerac F0 broodstock. We considered panmixia, thus the mating plan was one factorial design with all males and females from a given brood year. As the sex of trapped and released fish was unknown, a random arbitrary sex was assigned to each of them to achieve a balanced sex ratio.
Mating plans used to simulate salmon offspring from years of birth 2010–2014 in the Garonne-Dordogne basin restocking program.
2.3 Parentage assignment
All 4000 simulated individuals from a given salmon run (1000 wild-born and 1000 per hatchery) were assigned using Accurassign, a likelihood-based parentage assignment software (Boichard et al., 2014), with 10.000 simulations to set up assignment thresholds. Missing genotype rate was set to 1%, close to the observed value of 1.16% in the genotypes database, and genotyping error rate was set to 1%. According to Boichard et al. (2014), genotyping error rate is not a key parameter in their algorithm, and has to be low enough to penalize mismatches, but not too low to avoid exclusion based on a single marker incompatibility, and 1% is the default value. For the salmon run in year N, potential parents against which individual genotypes were tested included all hatchery broodstock used in years N-2 to N-6. This was done to have the same mating plan as the one used to analyse real returning salmon, for which the possibility that a juvenile may stay up to three years in fresh water is considered. However, simulated genotypes were only from parents in years N-2 to N-4, as the vast majority of salmon is expected to stay only one year in freshwater. To mimic reality, the genotypes of the parents of the simulated wild-born offspring were not included in the parent genotype database. The total number of potential parents was 5093 (3274 ♀, 1819 ♂) for the 2014 run, 5584 (3532 ♀, 2052 ♂) for the 2015 run and 5796 (3643 ♀, 2153 ♂) for the 2016 run.
Only offspring and parents which had a minimum of six properly genotyped loci out of the nine were included in the analysis, the other were qualified as non-compliant (NC).
Fish were assigned to their parents solely based on their genotype, and mating plan information was used only a posteriori to classify assignments as follows:
-
Assigned within mating plan (AssW) when the software assigned the individual to a single parental pair, which was part of the recorded mating plan
-
Assigned out of mating plan (AssO) when the software assigned the individual to a single parental pair, which was out of the recorded mating plan
-
Polyassigned (Poly) when two or more parent pairs were compatible with the offspring, but likelihood differences did not permit to rank them with sufficient confidence
-
Not assigned (Nass) when no parent pair was compatible with the offspring.
Given that the true parent pair was known for all simulated hatchery offspring, all assignments could be qualified as true or false.
Parentage assignment was also carried out for all returning individuals sampled at fish traps in 2014 to 2016 salmon runs following same approach (i.e. using the same parental genotype data set and the same mating plans).
Finally, in order to assess the reliability of the recorded mating plans, F1 individuals from the F1 hatcheries were assigned to their F0 parents from Bergerac, from years of birth 2008 to 2014.
2.4 Statistical analysis
Our aim was to estimate the true number of wild-born individuals among returning fish in a given salmon run, using assignment results from both the returning and simulated individuals, as well as to evaluate the reliability of assigning an individual fish to a “hatchery” or “wild-born” origin, depending on parentage assignment results.
Parentage assignment results from simulated offspring were summarized as proportion of individuals assigned within the mating plan P (AssW) and proportion of individuals with other assignment results P (other), which included all results (AssO, Poly, Nass) other than AssW.
From simulated hatchery fish, we could estimate P (other), conditional on the fact that animals were from hatchery origin, which was noted P (other|hatch). Similarly, from simulated wild-born fish, we could estimate P (other|wild). This was done for each simulated salmon run from 2014 to 2016.
The proportion of individuals with other assignment results in real data P (other) was estimated from the returning individuals of each salmon run from 2014 to 2016. Using Bayes'theorem, we could derive the probability of being wild for a returning individual, conditional on being assigned as “other”: (1)
(1)
Similarly, we could derive the probability of being from hatchery origin, conditional on being assigned as “other”: (2)
(2)
Similar formulae were derived for: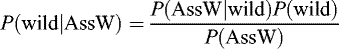 (3)
(3)
 (4)
(4)
Since (5)
(5)
and (6)
(6)
Equations (1), (2), (5) and (6) can be combined to obtain an estimate of the proportion of wild-born individuals in a given salmon run: (7)
(7)
This proportion P(wild) of wild-born returning salmon was estimated for the 2014 to 2016 salmon runs. This estimate may be modified if non-compliant parents are excluded from the analysis because they have only six or less loci genotyped (or have not been sampled). If P(NC) is the proportion of non-compliant hatchery parents, a reasonable hypothesis is that a proportion P(NC) of the hatchery offspring will be identified as wild (i.e. from unknown parents). If P
′(wild) and P
′(hatch) are the proportions of wild-born and hatchery fish taking into account the fact this proportion of non-compliant parents, then (8)
(8)
As P
′(hatch) + P
′(wild) = 1 (Eq. (7)) and thus P
′(hatch) = 1 − P
′(wild), equation (6) can be re-arranged as: (9)
(9)
3 Results
Globally, the parentage assignment procedure was very accurate in all simulated salmon runs ( Tab. 4). These formulae are implemented in the spreadsheet provided as Supplementary Material 1. The vast majority of hatchery-born simulated offspring was assigned within the mating plan (96.7%), contrary to wild-born simulated offspring (2.7%). However, assignment success was not symmetrical for non-assigned fish, which were 0% of the hatchery simulated offspring, but only 16.1% of the wild-born simulated offspring. Indeed, the most represented category among wild-born simulated offspring was poly-assigned fish (58.3%) followed by fish assigned out of the mating plan (22.9%). When assignment results were grouped in the “other than assigned within the mating plan” category, there was a clear differentiation between hatchery and wild-born simulated fish, with 3.3% of hatchery fish and 97.3% of wild-born fish classified as “other”.
Real returning salmon were assigned in significant numbers both to the AssW category (typical of hatchery simulated salmon) and to the “other than AssW” category (typical of wild-born simulated salmon), showing that these returning fish were a mixture of wild-born and hatchery salmon. Using equation (7), we could estimate the proportion of wild salmon in the 2014, 2015 and 2016 salmon runs ( Tab. 5), which was 32.1% on average. Taking into account the proportion of non-compliant parents in the reference mating plans for the different runs, which was 1.8% for 2014, 1.7% for 2016 and 2.2% for 2016, the corrected proportion was 30.7% wild-born fish on average.
The probability that a given individual was from hatchery origin if it was assigned within the mating plan (AssW) was very high, 98.7% on average. The probability that a given individual was wild-born if assigned as “other” (AssO, Poly, NAss) was also very high (93.3% on average).
Assignment rates of F1 hatchery individuals to their F0 parents were high, 95.8% on average, with 93.9% assigned within the mating plan and 1.9% assigned out of the mating plan ( Tab. 6). Assignment out of the mating plan was rather variable, 0.5% or lower in four years, 2.4% in 2009, 4.7% in 2013 and 4.3% in 2014. Poly assignments were more stable across years, around 0.6%. Unassigned offspring were 3.6% on average.
Parentage assignment of Atlantic salmon offspring with simulated genotypes at 9 microsatellite markers, for three salmon run years (2014–2016) in the Garonne-Dordogne Basin.
Probabilities associated to the 2014–2016 salmon runs (real data) in the Garonne-Dordogne basin.
Assignment of F1 salmon broodstock to their F0 parents.
4 Discussion
We showed that the parentage tracing system (9 microsatellites, analysed with Accurassign) used to monitor the Garonne-Dordogne Atlantic salmon restocking program was highly efficient, as 96.7% of the simulated hatchery fish could be traced back to a single parental pair belonging to the mating plan, and among those, the right parent pair was identified in 99.9% of cases (Tab. 4). This was true, despite the very large number of potential parents tested, which was higher than 5000 in all cases. For the 2016 salmon run, there were 5796 parents (3643 ♀, 2153 ♂) which corresponds to 7843379 potential families, considering the fact that the mating plan was not used in the assignment procedure per se, but only a posteriori to differentiate animals that were assigned within or outside of the mating plan. This is an excellent result, which is in line with those obtained in other salmonid restocking programmes (Steele et al., 2019). For example, 91.6–94.8% assignment rates were obtained by Beacham et al. (2019) in the coho salmon (Oncorhynchus kisutch) program of British Columbia with 304 SNPs genotyped. Interestingly, in the present study, the observed assignment rate (96.7%) was higher than the theoretical exclusion power (90.2%) that we estimated in Material and Methods for the marker set used, in a design with 5500 potential parents (3500 ♀, 2000 ♂), using the formula from Vandeputte (2012). This is most likely due to the fact that Accurassing uses a maximum likelihood algorithm, which is more efficient than simple exclusion (Boichard et al., 2014).
Not unexpectedly, we showed that data were at first sight more difficult to interpret for wild-born simulated animals. By construction, the parents of those wild-born fish were not present among the potential parents. Despite this, 25.6% of these wild-born fish were assigned to a single parental pair, 58.3% were assigned to multiple parental pairs, and only 16.1% were declared unassigned. It is not surprising that among the several millions of possible parental pairs, some present a likelihood to be compatible with the offspring that may cheat the assignment software. However, we could see that those wild-born individuals which were assigned to a single pair were spread across the full factorial mating scheme with all possible male-female combinations, and only a small proportion of them was within the effectively performed mating scheme. Only 2.7% of the wild fish were assigned to a single pair within the real mating scheme, and 22.9% were assigned out of the mating scheme. Thus, 89.4% of the wild fish that were assigned to a single pair were assigned to families that were not supposed to exist. This gave a rather efficient solution to identify these animals as not being of hatchery origin, especially as only 0.2% of the real hatchery fish were assigned to those “out of plan” families. Indeed, the real mating plan used to analyse the 2016 salmon run was composed of 28430 families out of a theoretical total of 7843 379 in 2016 (thus 0.4% of the total number of families). If families of the wrongly assigned wild fish were really randomly spread all across the full factorial mating plan, we would expect that 99.6% of them would be assigned as “out of plan” instead of 89.4%. It is probably due to the fact that many of the returning salmon used as parents for the wild simulated individuals are from hatchery origin, and therefore have genotypes that are closer to those of effectively used families than random crosses. Nevertheless, the classification remains highly efficient for discriminating wild-born from hatchery individuals (Tab. 5), and enabled us to estimate that an average 32.1% of the returning fish were wild-born in the three salmon runs studied. This estimate could be refined by taking into account the proportion of parental fish with missing genotype, which was 1.9% on average. The potential offspring of those fish could not be assigned to their parents, and thus were considered wild-born. When this issue was accounted for, the proportion of wild-born fish reduced to 30.7%. With this level of missing data, the consequences are limited, however this highlights the necessity to collect parental DNA with particular care, as it can lead to very high numbers of unassigned fish when parents sampling is incomplete (Araki et al., 2009). One more potential issue here is the fact that we assigned individuals as “wild-born” (implicitly from reproduction events in the Garonne-Dordogne river system) when they could not be assigned to Migado hatchery broodstock. An alternative explanation may be that those individuals were straying from other river systems. Indeed, it was shown in Southern France that up to 12–23% of returning salmon in the river Nivelle were from the nearby Bidasoa river population (Valiente et al., 2010). However, the distance between Nivelle and Bisadoa estuaries is very short (10 km), while in the case of Garonne-Dordogne, the closest salmon rivers are Loire (210 km to the North) and Adour (230 km to the South), which makes straying much more unlikely. Indeed, proven examples of recolonization by straying individuals from other river systems mostly imply nearby rivers: 7 km in Vasemägi et al. (2001) and Grandjean et al. (2009), mostly less than 60 km in Jonsson et al. (2003), but distant straying (>100 km), although less frequent, can also happen (Jonsson et al., 2003; Perrier et al., 2009). It is also suggested that straying salmon tend to stray more in unoccupied habitats than in rivers with an existing population (Vasemägi et al., 2001). Taken together, these observations suggest that while straying from other river systems cannot be ruled out, it is unlikely to represent the majority of the “wild-born” salmon identified here.
The efficiency of our approach to identify wild-born fish is also very much dependent on the exactness of the mating plan, which allowed to classify as “wild-born” those individuals which were assigned by the software to a parent pair that was not in hatchery records. However, if the mating plan was poorly recorded, individuals from families that were not recorded would have appeared as “assigned out of plan”, and thus as wild-born. We did not have data to evaluate the exactness of the mating plan of F2 individuals, but could do it for the F1. “Out of plan” assignments were 1.9% on average, showing that the recording system put in place was globally efficient in the Bergerac hatchery, and is thus likely to be equally efficient in the other hatcheries. We could see that “out of plan” assignments were very low (0.5% or less) in four of the years studied, corresponding to the expected values obtained in simulated offspring, for which the mating plan is exact by nature (BG sim in Tab. 4). However, these “out of plan” assignments reached significant values (2.4–4.7%) in three years. This is indicative that some mistakes happen in the recording of mating plans, albeit at low levels. Specifically, in 2013, 85 of the 90 “out of plan” offspring came from a single male, which was thus most probably participating, but was not recorded as such. In addition, we could see that unassigned individuals were more numerous (3.6% on average) than expected by simulation (0.0%, see BG sim in Tab. 4). This is likely due to the fact that F1 individuals from several years may coexist in F1 hatchery tanks, and that their DNA is sampled at the time of first reproduction, year of birth being assessed based on their size and maturity status. In a given year of DNA collection, it is thus likely that a few individuals are not from the alleged year of birth, which leads to their parents not being considered as potential parents in the analysis, and then to lack of assignment. However, this is not the case for the mating plans used to analyse migrants, where all potential parents are recorded. Nevertheless, if the power of the marker set was higher, the reliance on the mating plan could be minimized, as we would expect to see much less “assigned” (and poylassigned) individuals among wild-born ones, and many more “unassigned” ones. Therefore, since 2015, all individuals are genotyped for 17 microsatellite markers, but it will take several years before all potential parents of a given run have genotypes at 17 markers, thus the present approach remains useful, especially since we expect that even with more markers, the proportion of assigned and polyassigned fish within the wild ones will be strongly reduced but may not fall to zero.
5 Conclusion
We showed that in the context of the Garonne-Dordogne Atlantic salmon restocking programme, parentage tracing with microsatellite markers was efficient to discriminate hatchery-born from wild-born individuals when DNA samples of wild-born parents are not available. Practically speaking, we showed that individuals assigned within the known mating plan were from hatchery origin with 98.7% certainty. As traceability of the age and place of release of all mating plans is implemented in the recording system, this will enable the study of the most suitable sites and stages for restocking, including very young stages (eyed eggs, fry) at which physical tagging is not possible. In addition to those classical approaches, identifying wild-born animals, also with a high level of certainty (93.3%), will pave the way to studies on the abundance of those wild reproduction events, and on possible divergence between the wild-born and the hatchery individuals. It is of special importance to properly identify wild-born fish in such a restoration program, as establishing a self-sustained population is the final aim of the program. In this program, as the choice was made to stock mostly first-feeding fry, for logistic reasons, it is not possible to use adipose fin-clipping or coded wire tags to identify hatchery-born fish, and then, by difference, wild-born ones. Thus, demonstrating that they may be identified using genetic tagging, as we did in this study, is a key step to an efficient monitoring of the progress of the Migado program towards its objectives. A second potential benefit of the ability to identify wild-born individuals would be to use them (instead of randomly sampled migrants, most of which are presently of direct hatchery origin) as F0 parents in the Bergerac breeding center. This could be an interesting option to increase genetic diversity and counteract domestication selection in Migado hatcheries.
6 Data availability
Genotypes and mating plans available as: Vandeputte, Marc; Bestin, Anastasia; Fauchet, Louarn; Allamellou, Jean-Michel; Bosc, Stéphane; Menchi, Olivier; Haffray, Pierrick, 2020, “data for “Can we identify wild-born salmon from parentage assignment data? A case study in the Garonne-Dordogne rivers salmon restoration programme in France.””, https://doi.org/10.15454/VXSILB, Portail Data INRAE.
Supplementary Material
Workbook with macros to simulate salmon offspring genotypes and estimate the proportion of wild-born salmon. Access Supplementary Material
Acknowledgements
We wish to thank the personnel of the Migado hatcheries in Bergerac, Castels and Pont Crouzet and of the hatchery of the Fishermens' Federation of Hautes Pyrénées in Cauterêts for efficient management and data collection. We also thank Matthieu Chanseau from Office Français de la Biodiversité, Patrick Chèvre and René Guyomard from INRAE, for their initial contribution to the project. This work was financed by the “Programme général de restauration du saumon en Nouvelle Aquitaine” (European Union FEDER programme grant n° 2019–8015210, Agence de l'Eau Adour Garonne grant n°240.24.2120, Région Nouvelle Aquitaine grant n°EL8949220) and the “Programme de restauration des migrateurs du bassin de la Garonne en Occitanie” (European Union FEDER programme grant n° MP0025359, Agence de l'Eau Adour Garonne grant n°240.31.2335) as well as by Office Français de la Biodiversité.
Appendix A
Proportion of one sea winter, two sea winters and three sea winters salmon recorded in the 2014–2016 salmon runs in the Garonne-Dordogne river system.
| Sea age of salmon for each run | |||
|---|---|---|---|
|
|
|||
| 1SW (%) | 2SW (%) | 3SW (%) | |
| 2016 run | 43 | 53 | 3 |
| 2015 run | 16 | 78 | 6 |
| 2014 run | 9 | 87 | 4 |
References
- Araki H, Cooper B, Blouin MS. 2009. Carry-over effect of captive breeding reduces reproductive fitness of wild-born descendants in the wild. Biol Lett 5: 621–624. [PubMed] [Google Scholar]
- Aykanat T, Johnston SE, Cotter D, Cross TF, Poole R, Prodőhl PA, Reed T, Rogan G, McGinnity P, Primmer CR. 2014. Molecular pedigree reconstruction and estimation of evolutionary parameters in a wild Atlantic salmon river system with incomplete sampling: a power analysis. BMC Evol Biol 14: 68. [PubMed] [Google Scholar]
- Beacham TD, Wallace C, Jonsen K, McIntosh B, Candy JR, Willis D, Lynch C, Moore J-S., Bernatchez L, Withler RE. 2019. Comparison of coded-wire tagging with parentage-based tagging and genetic stock identification in a large-scale coho salmon fisheries application in British Columbia, Canada. Evol Appl 12: 230–254. [PubMed] [Google Scholar]
- Boichard D, Barbotte L, Genestout L. 2014. AccurAssign, software for accurate maximum-likelihood parentage assignment, in: Proceedings of the 10th World Congress on Genetics Applied to Livestock Production, p. 397. [Google Scholar]
- Evans ML, Johnson MA, Jacobson D, Wang J, Hogansen M, O'Malley KG. 2015. Evaluating a multi-generational reintroduction program for threatened salmon using genetic parentage analysis. Can J Fish Aquat Sci 73: 844–852. [Google Scholar]
- Grandjean F, Verne S, Cherbonnel C, Richard A. 2009. Fine-scale genetic structure of Atlantic salmon (Salmo salar) using microsatellite markers: effects of restocking and natural recolonization. Freshw Biol 54: 417–433. [Google Scholar]
- Griot R, Allal F, Brard‐Fudulea S, Morvezen R, Haffray P, Phocas F, Vandeputte M. 2020. APIS: An auto-adaptive parentage inference software that tolerates missing parents. Mol Ecol Resour 20: 579–590. [Google Scholar]
- Hess MA, Rabe CD, Vogel JL, Stephenson JJ, Nelson DD, Narum SR. 2012. Supportive breeding boosts natural population abundance with minimal negative impacts on fitness of a wild population of Chinook salmon. Mol Ecol 21: 5236–5250. [PubMed] [Google Scholar]
- Jamieson A. 1965. The genetics of transferrins in cattle. Heredity 20: 419–441. [PubMed] [Google Scholar]
- Jonsson B, Jonsson N, Hansen LP. 2003. Atlantic salmon straying from the River Imsa. J Fish Biol 62: 641–657. [Google Scholar]
- Kalinowski ST, Taper ML, Marshall TC. 2007. Revising how the computer program cervus accommodates genotyping error increases success in paternity assignment. Mol Ecol 16: 1099–1106. [CrossRef] [PubMed] [Google Scholar]
- Kuparinen A, Keith DM, Hutchings JA. 2014. Allee effect and the uncertainty of population recovery. Conserv Biol 28: 790–798. [PubMed] [Google Scholar]
- McGinnity P, Prodhöl P, Ferguson A, Hynes R, O'Maoileidigh N, Baker N, Cotter D, O'Hea B, Cooke D, Rogan G, Taggart J, Cross T. 2003. Fitness reduction and potential extinction of wild populations of Atlantic salmon, Salmo salar, as a result of interactions with escaped farm salmon. Proc R Soc B Biol Sci 270: 2443–2450. [Google Scholar]
- Paterson S, Piertney SB, Knox D, Gilbey J, Verspoor E. 2004. Characterization and PCR multiplexing of novel highly variable tetranucleotide Atlantic salmon (Salmo salar L.) microsatellites. Mol Ecol Notes 4: 160–162. [Google Scholar]
- Perrier CP, Evanno GE, Belliard JB, Guyomard RG, Baglinière J-LB-L. 2009. Natural recolonization of the Seine River by Atlantic salmon (Salmo salar) of multiple origins. Can J Fish Aquat Sci 67: 1–4. [Google Scholar]
- Slettan A, Olsaker I, Lie Ø. 1995. Atlantic salmon, Salmo salar, microsatellites at the SSOSL25, SSOSL85, SSOSL311, SSOSL417 loci. Anim Genet 26: 281–282. [CrossRef] [PubMed] [Google Scholar]
- Steele CA, Hess M, Narum S, Campbell M. 2019. Parentage‐based tagging: reviewing the implementation of a new tool for an old problem. Fisheries 44: 412–422. [Google Scholar]
- Stephens PA, Sutherland WJ. 1999. Consequences of the Allee effect for behaviour, ecology and conservation. Trends Ecol Evol 14: 401–405. [CrossRef] [PubMed] [Google Scholar]
- Thibault M. 1994. Aperçu historique sur l'évolution des captures et des stocks. In: Le Saumon Atlantique, Biologie et Gestion de La Ressource. Guegen, J.C. & Prouzet, P., Plouzané, Ifremer, pp. 173–183. [Google Scholar]
- Valiente AG, Beall E, Garcia‐Vazquez E. 2010. Population genetics of south European Atlantic salmon under global change. Global Change Biol 16: 36–47. [Google Scholar]
- Vandeputte M. 2012. An accurate formula to calculate exclusion power of marker sets in parentage assignment. Genet Sel Evol 44: 36. [Google Scholar]
- Vandeputte M, Haffray P. 2014. Parentage assignment with genomic markers: a major advance for understanding and exploiting genetic variation of quantitative traits in farmed aquatic animals. Front Genet 5: 432. [PubMed] [Google Scholar]
- Vasemägi A, Gross R, Paaver T, Kangur M, Nilsson J, Eriksson L-O. 2001. Identification of the origin of an Atlantic salmon (Salmo salar L.) population in a recently recolonized river in the Baltic Sea. Mol Ecol 10: 2877–2882. [PubMed] [Google Scholar]
- Wang J. 2012. Computationally efficient sibship and parentage assignment from multilocus marker data. Genetics 191: 183–194. [CrossRef] [PubMed] [Google Scholar]
Cite this article as: Vandeputte M, Bestin A, Fauchet L, Allamellou J-M, Bosc S, Menchi O, Haffray P. 2021. Can we identify wild-born salmon from parentage assignment data? A case study in the Garonne-Dordogne rivers salmon restoration programme in France. Aquat. Living Resour. 34: 7
All Tables
Types of assignments and errors in parentage assignment with genomic markers, depending on the availability of parental genotypes.
Variability and assignment power of the nine microsatellite markers estimated from the F0 salmon broodstock present in Bergerac hatchery from 2010 to 2014 (156 individuals).
Mating plans used to simulate salmon offspring from years of birth 2010–2014 in the Garonne-Dordogne basin restocking program.
Parentage assignment of Atlantic salmon offspring with simulated genotypes at 9 microsatellite markers, for three salmon run years (2014–2016) in the Garonne-Dordogne Basin.
Probabilities associated to the 2014–2016 salmon runs (real data) in the Garonne-Dordogne basin.
Current usage metrics show cumulative count of Article Views (full-text article views including HTML views, PDF and ePub downloads, according to the available data) and Abstracts Views on Vision4Press platform.
Data correspond to usage on the plateform after 2015. The current usage metrics is available 48-96 hours after online publication and is updated daily on week days.
Initial download of the metrics may take a while.


