| Issue |
Aquat. Living Resour.
Volume 35, 2022
|
|
|---|---|---|
| Article Number | 4 | |
| Number of page(s) | 14 | |
| DOI | https://doi.org/10.1051/alr/2022004 | |
| Published online | 24 May 2022 | |
Research Article
COI gene sequencing reveals genetic population structuring and a separate lineage of the green tiger prawn Penaeus semisulcatus in the Gulf of Suez and the Bitter Lakes, Egypt
1
Zoology Department, Faculty of Science, Menoufia University, Shebin El-Kom, Menoufia, Egypt
2
National Institute of Oceanography and Fisheries, NIOF, P.O. Box 371, Suez, Egypt
* Corresponding author: Khaled.mohammed@icman.csic.es; khaledmohammed@science.menofia.edu.eg.
Handling Editor: Roman Wenne
Received:
30
December
2021
Accepted:
7
March
2022
The green tiger prawn Penaeus semisulcatus is one of the major commercial trawl crustaceans in the Red Sea, the Arabian Gulf, and the Indian Ocean. Herein, we aimed to assess its genetic diversity and phylogeographic criteria in the North West (NW) of the Red Sea. Prawn samples were collected from the Bitter Lakes; the North; and the South of the Gulf of Suez in Egypt. The hypervariable, 5´ barcode area of the Cytochrome oxidase subunit 1 (COI) gene was PCR-amplified and sequenced in all samples. The resulting sequences were subjected to several genetic, phylogenetic, and population genetics' analyses. Consequently, clear signs of genetic populations’ diversity, expansion; and possible structuring were identified in different areas in the NW Red Sea. Furthermore, extremely discordant genetic connectivity was found between the populations around the Arabian Plate (i.e. of the Red Sea and Strait of Hormuz) and the IndoWest Pacific, resulting in characterizing the Arabian plate populations as a separate lineage of P. semisulcatus. Based on these findings, it is important to investigate whether there are other discriminative genomic, morphological and/or morphometric differences among the genetically identified lineages and populations. Consequently, differential conservational strategies of different populations of this species should be applied in the studied areas.
Key words: Arabian Plate / cryptic / genetic structuring / lineages / Penaeus semisulcatus / populations / Red Sea
© K. Mohammed-Geba and A. Yousif, published by EDP Sciences 2022
 This is an Open Access article distributed under the terms of the Creative Commons Attribution License (https://creativecommons.org/licenses/by/4.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
This is an Open Access article distributed under the terms of the Creative Commons Attribution License (https://creativecommons.org/licenses/by/4.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
1 Introduction
The family Penaeidae (first described by Rafinesque in 1815) comprises 226 species in 32 genera, 77% of which are located in the Indo-Pacific (Ditty, 2014). This is possibly related to the central Indo-Western Pacific origin of genus Penaeus, which passed through several eastward and westward migrations, founder effects, and speciation events, especially as a result of Pleistocene glaciations (Alam et al., 2017; Wang et al., 2020).
The green tiger prawn Penaeus semisulcatus was first described by De Haan in Japan in 1844. This prawn occupies a very wide area, from the South and West of the Pacific Ocean to the Eastern Mediterranean Sea (WoRMS, 2021). It contributes major economic benefits for fisheries of the Pacific and the Indo-West Pacific (IWP) areas and their related habitats (Dore and Frimodt, 1987). Also, its capture is experiencing a continuous rise by countries of the Red Sea, mainly Egypt and Saudi Arabia (Rabaoui et al., 2017; El-Ganainy and Yassien, 2021). In Egypt, the Gulf of Suez is located at the North of the Red Sea, connecting it with the Mediterranean Sea through Suez Canal that was dug in the 19th century. Along the Canal trajectory, shallow small lakes are connected to the canal basin, known as Bitter lakes. Penaeus semisulcatus and P. (Marsupenaeus) japonicus Spence Bate, 1888 constitute together the main category of large shrimps in the main port of fisheries landings, i.e. the Attaka Port, in Suez (Yousif, 2003). This category represents around 12% of the total seasonal trawl catch in the Gulf. The seasonal catch of large shrimps deeply fluctuates and increased in the last three seasons (Fig. 1). Penaeus semisulcatus is also well known as a Lessepssian migrant. It crossed the Suez Canal and colonized the coasts of the eastern and north-eastern Mediterranean, especially preferring soft-silt bottoms (Boudouresque, 1999, Ragonese et al., 2011). It reached so far to the English Channel, being its first record in that area was in Le Havre Harbour, Normandy, France in 2016 (Pezy et al., 2017).
Several aspects of P. semisulcatus biology exhibit contradictions among its different populations. For example, in the Southwestern part of the Arabian Gulf, its spawning peaks are in summer and winter (Rabaoui et al., 2017), while in north-eastern waters of the Gulf its spawning peaks are in the early winter and early spring (Niamaimandi et al., 2008). Also, salinity tolerance among different populations of this species varies considerably, between 20 and 40 practical salinity units (PSU) (Soyel and Kumlu, 2003). More recently, molecular clues proved that there are several genetic lineages of P. semisulcatus in the Arabian Gulf (Jahromi et al., 2019).
Owing to its economic importance, P. semisulcatus was genetically investigated in the Gulf of Suez and through the Suez Canal in several works before. For example, Sharawy et al. (2016, 2017) carried out different genetic identification for this species using several molecular markers, including cytochrome oxidase subunit 1 (COI), 16S rDNA, and 18S rDNA sequencing. These works are of the pioneer efforts for genetic characterization of this species in the Northern part of the Red Sea and the South East of the Mediterranean Sea. However, very scarce knowledge is available about the genetic populations’ structuring of this species in different areas of the Red Sea. Similarly, the relationship between P. semisulcatus lineage in the Red Sea-Mediterranean Sea link with other lineages of this species in the Indian Ocean lacks also detailed genetic analyses.
The present research work aimed to resolve three key issues about the genetic diversity of this species in the North of the Red Sea. First, to assess the genetic diversity of this species in the Gulf of Suez and the Bitter Lakes, where the most abundant fisheries of this species in Egypt exist. Second, to test whether there is any population structuring of P. semisulcatus in these two areas. Third, to identify the exact genetic lineages to which Red Sea P. semisulcatus belong, which is an almost unstudied aspect. Therefore, the present study was designed to provide direct genetic data regarding the presence of characteristic populations in the Northern Red Sea P. semisulcatus, as well as to determine the phylogeographic location of this species among the other populations of it in the world.
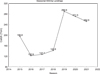 |
Fig. 1 The annual catch of ¨large shrimps¨ category, i.e. Penaeus semisulcatus and Penaeus japonicus in the Gulf of Suez, Egypt. Total numbers of catch in the period between 2014-2022 are shown in tons. Source: General Authority for fisheries resources development in Egypt (GAFRD), Suez branch (2022). |
2 Materials and methods
2.1 Collection of samples
Ninety fresh green tiger prawns (Penaeus semisulcatus) samples were collected by an artisanal boat from the Bitter Lakes (BL) (N30º26′61.7′′, E32º 44′98.6′′) and by trawling from two different locations in the Gulf of Suez, i.e. Sadat (SA), at the North of the Gulf: N29º45′74′′, E32º30′′17′; and Abu Regm (AR), at the South of the Gulf: N28º43′′35′, E32º51′′13′, as 30 samples (n = 30) from each site (Fig. 2). Approximately, 100 mg of each shrimp muscles’ tissue were preserved in separate 1.5 mL sterile tubes containing 96% ethanol and stored at –20 °C. Ethanol-preserved shrimp samples were then transferred to the Molecular Biology and Biotechnology Laboratory of the Zoology Department in the Faculty of Science of Menoufia University in Egypt for subsequent genetic analyses.
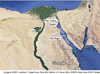 |
Fig. 2 Sampling locations, shown as white points, for Penaeus semisulcatus from Bitter Lakes (N30º26'61.7'', E32 º 44'98.6'') and Gulf of Suez [Sadat (N29º45'74", E32º30'17) and Abo Regm (N28º43'35", E32º51'13)] at the North-Western Red Sea, Egypt. Photo from GoogleMapsTM (map credits are shown below the photo). |
2.2 DNA purification and PCR-amplification for COI gene
Total DNA was extracted from 10 mg of the pleonal musculature of each shrimp sample using Chelex® 100 sodium form (Walsh et al., 1991). The universal primers LCOI490 (5′-GGTCAACAAATCATAAAGATATTGG-3′) and HCO2198 (5′-TAAACTTCAGGGTGACCAAAAAATCA-3′) (Folmer et al., 1994) were used for the amplification of an approximately 660 base pair (bp) fragment from the 5′ end of the mitochondrial cytochrome oxidase subunit 1 (COI) barcode of life gene. The amplification reactions were performed using the same conditions and program as in Folmer et al. (1994). The PCR products were electrophoresed in a 1% agarose gel. The correct-sized PCR products were sent to Macrogen Inc. (Seoul, South Korea) for purification and sequencing using the conventional Sanger chain termination method, applying the same-sequence primers used for PCRs for forward and reverse sequencing for each PCR product.
2.3 DNA sequences and population genetics’ analyses
COI sequences’ chromatograms were manually checked and trimmed to remove non-informative nucleotide sites at both extremities, using the freeware Chromas Lite 2.1. Each sequence was individually compared to GenBank database using BLAST algorithm (http://blast.ncbi.nlm.nih.gov/Blast.cgi) and IDS (https://www.boldsystems.org/index.php/IDS_OpenIdEngine) to BOLD. All sequences were then aligned using CLUSTAL W Multiple alignment in Mega11 platform (Tamura et al., 2021). Penaeus semisulcatus COI sequences were translated into the primary amino acids’ sequences using invertebrate mitochondrial codons in Mega11 to confirm the absence of stop codons that characterize the nuclear copies of mitochondrial genes (NuMTs). Later on, all the obtained and checked P. semisulcatus COI sequences were deposited in the GenBank database under the accession numbers (acc. No. OM063021 - OM063096).
The sequences sets were later defined using DnaSP 6 Software (Rozas et al., 2017) based on their pertinence to each of the three sampling locations (Bitter Lakes, Sadat, Abo Regm). Furthermore, COI gene haplotypes existing in common and separately among the three sampling locations were determined, together with haplotype diversity and nucleotide diversity indices. Moreover, the same sets-defined alignment was utilized to estimate populations’ expansions and raggedness as inferred from the abundance of singleton haplotypes and the homogeneity of nucleotide pairwise distances there, using R2 parameter (Ramos-Onsins and Rozas, 2002) and the index of raggedness (r) (Harpending, 1994), respectively. Mismatch distribution of pairwise distances was calculated and plotted using DNAsp6 also, to further test whether the population of P. semisulcatus is under demographic equilibrium (i.e. multimodal curves) or undergoing recent population expansion (unimodal curve). Later on, the Egyptian haplotypes determined through DnaSP 6 Software were uploaded to the program PopArt 1.7. (http://popart.otago.ac.nz/index.shtml) to draw a median-joining haplotype network for demonstrating the interrelationships among different haplotypes. Furthermore, a universal network was drawn to demonstrate the interrelationships between P. semisulcatus haplotypes from around the Arabian Plate and the Indo-West Pacific countries. This new MJ network was based on the world haplotypes’ alignment file that will be mentioned in Section 2.5. herein.
Population sets were uploaded to the software ARLEQUIN 3.5.1.1 (Excoffier and Lischer, 2010) calculating the fixation index (Fst). Then, the matrix of Fst values calculated through ARLEQUIN was applied for performing Analyses of Molecular Variance (AMOVA) (Excoffier, 1992). Through AMOVA, Non-significantly different populations were grouped to exhibit the structure that shows the least within-population difference and the most among-populations difference. This, in the case of having different allelic frequencies in some of the groups in this species population, may refer to the existence of different populations. Finally, ARLEQUIN was utilized to estimate Tajima’s D and Fu’s Fs statistics (Tajima, 1989; Fu, 1997) that assess the neutral evolution.
2.4 Phylogenetic analyses
In order to analyze the phylogenetic inference of the Egyptian population of P. semisulcatus assayed in the present study within the world populations of the same species, COI sequences for P. semisulcatus were retrieved from GenBank and BOLD databases. These sequences belonged to the Indo-West Pacific region, the Arabian Gulf, the Red Sea, and the Mediterranean Sea. Accession numbers for these sequences are given in Table 1. After the removal of COI sequences that could not be aligned with our sequences, these sequences were merged into haplotypes using DnaSP 6 Software. Furthermore, a partial COI barcode sequence from the southern horseshoe crab Tachypleus gigas was retrieved from the GenBank (acc. No. JF896114.1) to serve as a tree outgroup. All sequences were aligned using CLUSTAL W in the Mega11.
The alignment of all haplotypes of P. semisulcatus, together with the outgroup, was uploaded to MrBayes 3.2.1 software (Ronquist et al., 2012), to confirm the identity of our Egyptian species using Bayesian inference (BI) for phylogenetic relationships. Four Markov Chains Monte Carlo (MCMC) chains were analyzed for 10 million (ngen = 10,000,000) generations, saving a tree each 1000 generations. The subsequent analyses were carried out after assuring an average standard deviation of split frequencies below 0.001. The number of burn-ins was identified using Tracer 1.7 (Rambaut et al., 2018). Tracer 1.7 was applied to identify the percentage of burn-ins to be discarded. This information was transferred to MrBayes 3.2.1. for constructing the trees. The result was then uploaded to TreeAnnotator software V. 1.8.3 for summarizing the information retrieved from tree samples, then the tree was viewed using the Interactive Tree of Life online algorithm (iTOL: https://itol.embl.de/).
To construct a Maximum Likelihood (ML) phylogenetic tree, the alignment file of P. semisulcatus haplotypes and the outgroup was saved as .fas (fasta) format, then exported to JModelTest software V. 2.1.10 (Darriba et al., 2012) for detection of best nucleotide substitution model. Then, the Fasta file was uploaded to Beauti software V. 1.8.3, with the substitution model determined using JModelTest, and 10,000,000 Markov chains were selected. The program was run once with this number of Markov chains, then two other runs were carried out, one with 50,000,000 chains and another with 100,000,000 chains. The resulting .xml files were opened using BEAST software V. 1.8.3 for estimating tree topologies. The resulting .log files from the three trials were uploaded to the program Tracer v1.6 to assess the quality of the results. The resulting trees were combined using LogCombiner software V. 1.8.3, then uploaded to TreeAnnotator software V. 1.8.3 for summarizing the information retrieved from tree samples produced through BEAST. The resulting consensus tree was then finally visualized using iTOL algorithm.
Accession numbers and locations for GenBank and BOLD-retrieved COI sequences of P. semisulcatus from different areas of the world.
2.5 Pairwise distances and species delimitation
The alignment created for different Red Sea, Arabian Gulf, and Indo-West Pacific (IWP) haplotypes, i.e. the resulting unique COI sequences’ data set, was analyzed using Kimura’s 2-parameter model (K2P). This aimed to estimate the mean within-population pairwise sequence divergences for all populations, using the procedure implemented by Witt et al. (2012) for the identification of species screening threshold (SST). The populations analyzed through this approach were separated into two main groups. The first group in the population included the Arabian Plate’s haplotypes (i.e. Northern Red Sea + Western part of the Strait of Hormuz). The second group included the IWP’s (Bangladesh + Sri Lanka + Malaysia) haplotypes. A ratio of more than 10 between the averages of these two populations’ pairwise distances was applied to delimit possible cryptic species. Furthermore, the average number of nucleotide substitutions per site (k) for different populations was identified using DnaSP6 software, to test the proximity of the resulting ratio to the Crustaceans’ species delimitation threshold of 0.16 substitutions per site, according to Lefebure et al. (2006) and Alam et al. (2016).
3 Results
3.1 DNA polymorphism in the COI barcode fragment
After manual trimming of non-informative initial and final nucleotide peaks from all sequences, a COI fragment length of ca. 586 nucleotides could be obtained in the nucleotide sequences alignment. The total number of polymorphic nucleotide sites was 20, characterizing 23 different haplotypes in the three sampled locations, i.e., Bitter Lakes, Sadat, and Abo Regm. Eight haplotypes were found in Bitter Lakes (BL), whereas 9 haplotypes were present in Sadat area (SA), and the other 12 ones were found in Abo Regm area (AR) (Tab. 2). Site-specific or private haplotypes were distributed as 4 for BL, 7 for SA, and 7 for AR.
Comparison of the 23 identified Egyptian haplotypes resulted in 99–100% sequence identity with a few P. semisulcatus samples that had acc. No LC175221, LC155214, LC155215 KR261586, and KR261588. However, much lower COI sequences’ identities (86–96%) were found with P. semisulcatus samples collected from more Southeastern areas of the Indian Ocean; i.e. KX585464–KX585491, KX585492–KX585624, and MT178728- MT178733.
Different haplotype diversity, nucleotide diversity, neutrality, and raggedness analyses values for Egyptian P. semisulcatus sampled from the Bitter Lakes and Gulf of Suez, Egypt.
3.2 Demographic estimators
Regarding the genetic-demographic estimators, all populations of the three areas, i.e. BL, SA, and AR exhibited significant negative values for Fu’s Fs and Tajima’s D neutrality tests. In accordance, the mismatch curve exhibited a unimodal pattern (Fig. 3). In addition, non-significant raggedness and low R2 values have resulted in the three analyzed groups, yet being that of BL the one with the highest raggedness index value among the three. Nucleotide diversities (π) were low in all locations, ranging between 0.001 and 0.002 (Tab. 2).
The constructed median-joining network for the identified 23 haplotypes (Hap) in the three populations assumed a star shape (Fig. 4), with a single main haplotype (Hap 1) that was found in the three locations. 17 samples from BL, 15 samples from SA, and 14 samples from AR belonged to Hap 1-the haplotypes that appear as the largest diameter circle in Figure 4. Few other haplotypes were shared among the sampling areas. These were Hap2, which was present in both BL and AR (BL + AR), Hap 5 (BL + AR), Hap 7 (BL + AR), and Hap 15 (SA + AR). Singleton haplotypes (i.e. private ones) were present in the three studied locations, ranging in percentage from 50% of total haplotypes present in BL, to about. 80% of total haplotypes are present in SA area. The singleton haplotypes in SA and AR slightly outnumbered these in BL (Tab. 2, Fig. 4). For the world haplotype network, two main groups were identified; one included the Arabian Plate populations, and the other included all the Indo-West Pacific populations (Fig. 5).
Regarding the genetic flow among the P. semisulcatus Red Sea populations analyzed herein, significant fixation could be identified between the populations of Bitter Lakes and Abo Regm area (Fst = 0.03, P < 0.05). However, Bitter Lakes and Sadat did not show fixation (Fst = 0.01, P > 0.05). Sadat and Abo Regm population showed also no fixation (Fst = 0, P > 0.05). Analysis of Molecular variance (AMOVA) carried out between different combinations of the three areas retrieved that the structure that gave the highest (Among populations) variations and the least (Within populations) variation was grouping Sadat and Abo Regm areas together, against Bitter lakes in a separate group (Tab. 3).
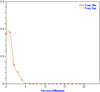 |
Fig. 3 Mismatch distribution for COI-based pairwise distances of the North-Western Red Sea and Bitter lakes’ populations of P. semisulcatus. |
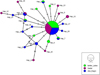 |
Fig. 4 Median-joining haplotype network for COI sequences in P. semisulcatus obtained from different areas in the North-West of the Red Sea. The branch length is proportional to the number of substitutions. Black transverse bars in the branches represent the polymorphic sites that differ among haplotypes. Circles represent haplotypes and their diameters are proportional to the haplotype frequencies. The Colour legend is shown on the down-right side of the graphic. |
 |
Fig. 5 Median-joining haplotype network for COI sequences in P. semisulcatus from the Red Sea, the Arabian Gulf, and the Indo-West Pacific region (Bangladesh, Sri Lanka, Malaysia). The colour legend is shown on the down-right side of the graphic. |
Analysis of molecular variance (AMOVA) of Fst values for grouped P. semisulcatus COI sequences from different sampling locations in the NW of the Red Sea.
3.3 Phylogenetic analysis
Both BI and ML trees exhibited the presence of two distinct phylogenetic clades encompassing all haplotypes analyzed in the present study. A single clade included all Southern and Southwestern Asian haplotypes of this species, i.e., of Malaysia, Sri Lanka, and Bangladesh (Figs. 6 and 7). Another separate clade included all P. semisulcatus samples from around the Arabian Tectonic Plate, i.e., the samples that were collected and genetically analyzed in the present study, and the other samples for the same species whose sequences were retrieved from the GenBank from the Arabian Gulf and Suez Canal, besides the ones from the Mediterranean coasts of Alexandria, Egypt. Posterior probabilities for these two phylogenetically sister clades were 100% in both BI and ML trees. Despite both trees exhibited similar topologies, the BI tree elucidated the presences of different subclades in both the Southern and Northern clades (Fig. 6).
 |
Fig. 6 BI phylogenetic analysis for the samples collected in the present study. Posterior probabilities are shown above the branches. Green highlighted: Gulf of Suez P. semisulcatus haplotypes (Hap), and the closely related ones form the Arabian Gulf and the Mediterranean Sea. Red-highlighted: Indo-West Pacific (IWP) haplotypes. Tree outgroup: the Southern Horseshoe crab Tachypleus gigas. |
 |
Fig. 7 Maximum-Likelihood (ML) phylogenetic tree for P. semisulcatus world haplotypes (Hap). Posterior probabilities are shown above the branches. Green highlighted: Gulf of Suez P. semisulcatus haplotypes (Hap), and the closely related ones form the Arabian Gulf and the Mediterranean Sea. Red-highlighted: Indo-West Pacific (IWP) haplotypes. Tree outgroup: the Southern Horseshoe crab Tachypleus gigas. |
3.4 Pairwise distances and species delimitation
COI-based genetic pairwise distances exhibited a clear disparity between the Arabian Plate haplotypes (Egypt and Iran), and the ones from Indo-West Pacific oceanic regions (Bangladesh, Sri Lanka, and Malaysia) (Fig. 8). The average pairwise genetic distances among COI fragments of the Indo-West Pacific ranged between 0.247 and 0.314, with an average of 0.284. However, the genetic pairwise distances among the Red Sea and Strait of Hormuz samples were between 0 and 0.011, with an average of 0.004. The calculated ratio of SST was 64.1, which is 6.4 times the species-screening-threshold (10x) for COI gene. Furthermore, the average number of nucleotide substitutions per site was 7.005 for the Indo-West Pacific oceanic populations, and 2.440 for the Arabian Plate (i.e. the Red Sea and the West of the Strait of Hormuz) populations, resulting in a ratio of 0.35, which exceed the double of the species delimitation value based on average nucleotide substitution rates.
 |
Fig. 8 Distribution of genetic pairwise distances among Gulf of Suez, Arabian Gulf, and Indo-West Pacific P. semisulcatus COI haplotypes in comparison to those in the GenBank and BOLD databases. |
4 Discussion
To the best of the authors’ knowledge, this is one of very few works that indicated the presence of some population structuring of P. semisulcatus in the Red Sea, yet the first to apply COI gene sequencing for assessing this diversity and structure. Despite this study can be related to the previous works of Sharawy et al., 2016, 2017 who carried out genetic identification for this species in the Red Sea, the present work provided new data about P. semisulcatus genetic structuring in the Gulf of Suez and Bitter Lakes, as well as connecting it to its Indian Ocean counterparts. This data can be of specific importance as this species is a major contributor to the bottom trawl fisheries in several areas in Egypt, especially the North West (NW) of the Red Sea in Egypt, as well as in the Sea of Oman and the Arabian Gulf (Yousif, 2003; Rabaoui et al., 2017; El-Aiatt et al., 2019). Despite being widely distributed in the continental waters of Eastern and Northern Africa, after entering the Mediterranean as a Lessepsian migrant; South and South East; and Australia; still many aspects in this prawn’s genetic diversity and populations’ structures are to be identified. Herein, our results indicated the presence of signs of population structuring of P. semisulcatus in the NW Red Sea.
In the present study, P. semisulcatus population structuring was revealed in the NW of the Red Sea, as indicated from the significant fixation between the Northern-most, i.e. Bitter Lakes, and the Southern-most, i.e. Abo Regm, locations. Besides, AMOVA showed that Bitter Lakes and Sadat area can be considered as a single population, while that of Abo Regm is another population. Furthermore, the location of Bitter Lakes exhibited the highest raggedness and the lowest expansion (as identified from the raggedness index and Tajima’s D statistic, respectively) among all the three locations, yet being all locations with low raggedness and significant recent expansion. These special criteria in the Bitter Lakes group can be expected due to the hypersalinity in that area, which still represents a globally-identified barrier for Red Sea marine species north-ward migrations (Bernardi et al., 2016). The capability of organisms to adapt to that high salinity barrier, besides the low salinity barrier of the Nile estuary, deeply affected the capability of Red Sea organisms to spread to the Mediterranean (Kiparissis et al., 2018). No literature could be found regarding the genetic diversity of prawn species within the Bitter Lakes, compared to the same species before and after this barrier. However, some international examples for clear genetic differences as a result of the presence of a different salinity barrier within the native ranges of prawn species are known. For example, freshwater outflow from the Yangtze River was related to the significant genetic differentiation between the East China Sea and the Yellow Sea populations of both cocktail shrimp Trachysalambria (Trachypenaeus) curvirostris and mantis shrimp Oratosquilla oratoria (De Haan, 1844) (Han et al., 2015; Du et al., 2016). Future works regarding the genetic diversity within P. semisulcatus in other areas, such as to the North of Bitter Lakes and in the Mediterranean, are required to identify the status of populations of this species along the Suez Canal and after.
Furthermore, the population of P. semisulcatus in the North-Western part of the Red Sea exhibited high percentages of haplotype diversities (ca. 50–70%). This exceeded the known for this species in other areas in the world using COI gene sequencing, i.e. in the Indo-West Pacific (Alam et al., 2017). However, much lesser nucleotide diversities were identified in the three assayed regions in NW of the Red Sea, in comparison to those found before in those in the IWP. High haplotype diversity; low nucleotide diversity; and negativity of Fu’s Fs and Tajima’s D statistics, which serve to denote the deviation from neutrality; can strongly indicate the presence of a large population of P. semisulcatus in the studied areas. This hypothesized large-sized population was subjected to recent population expansion, possibly after a period of low effective population size or bottleneck. This allows the retention of new alleles in the population but without enough time for the accumulation of more nucleotide substitutions among haplotypes (Grant and Bowen, 1998; Kasim et al., 2020). This recent population expansion can be further supported by the star-shaped haplotype network and the unimodal mismatch distribution curve of the pairwise distances. Similar conditions were found in several shellfish and finfish species. For example, high haplotype diversity, low nucleotide diversity, unimodal mismatch distribution were found in the Indo-West Pacific, overexploited yellow drum Nibea albiflora (Richardson, 1846) (Hd = 0.580–0.815, π = 0.00125–0.00219) (Xu et al., 2012). Furthermore, the cocktail shrimp T. curvisrostris exhibited moderate-high haplotype diversity and low nucleotide diversity (Hd = 0.6888 ± 0.0432, π = 0.0069 ± 0.0038) in the regions of the East China Sea and the Yellow Sea (Han et al., 2015). This pattern was attributed in that species to population range expansion, possibly before 103–109 thousand years ago (KYA), i.e., in the last interglaciation (Han et al., 2015). Also, the shovel-nosed lobster Thenus unimaculatus Burton and Davie, 2007 exhibited similar patterns in the Eastern Indo-West Pacific areas as those found of P. semisulcatus in the present study, which was explained by Saher et al. (2021) as signs of recent population expansion. However, other scenarios exist for this pattern of nucleotide and haplotype diversities, including the subjection of species to overexploitation and habitat destruction. This was suggested for several other species of shrimps around the world, including for example the pink shrimp Farfantepenaeus paulensis (Pérez Farfante, 1967) in the south-southeastern coast of Brazil (Teodoro et al., 2020) and the Indian prawn Fenneropenaeus indicus (H. Milne Edwards, 1837) in the Tanzanian coastal waters (Mwakosya et al., 2018). In all cases, these may represent signs of the necessity of more serious investigation of species state of conservation and application of through stock surveillance management strategies.
Another key aspect that could be identified in the present study was the discordant genetic patterns between the NW Red Sea and Strait of Hormuz populations of P. semisulcatus in one side from these in the IWP region in another. The presence of separate lineages within P. semisulcatus is known, yet being relatively recent. Through mitochondrial control region, 16S rDNA, and COI sequencing, it was identified that there are distinct mitochondrial DNA lineages, one in the Indian Ocean, represented by Malaysia, Sri Lanka, and Bangladesh; and another in the Western Pacific Ocean, represented by China (Alam et al., 2017). More recently, a boundary between the Indian and Pacific Ocean populations of P. semisulcatus could be identified in the strait of Malaca, separating the Indian and Pacific populations as two separate phylogenetic lineages (Halim et al., 2021). Yet, it was not possible to compare the results of this study with the present one owing to the different primers and barcoding areas applied between them. In the study of Alam et al. (2017), an SST ratio of 12.9 and substitution rate per site of 0.16 were found between the two lineages, and both were considered as clues for elucidating two cryptic species in the assessed region under the generic name P. semisulcatus. Cryptic species were suggested at an SST ratio of >10 (Witt et al., 2012). Also, a threshold ratio of 0.16 substitutions per site of the COI gene in Crustaceans was considered valid for species delimitation (Lefebure et al., 2006). Despite many Iranian COI sequences available at GenBank were not alignable with our ones, in contrast to almost all other ones from the IWP region, two sequences from the Strait of Hormuz could be successfully aligned. These were the ones that perfectly claded with the NW Red Sea samples analyzed in the present study. The presence of unique sequences of P. semisulcatus that possibly point to an unknown lineage was noted by Alam et al. (2017), yet their origin could not be identified. Also, two genetically and morphologically distinct forms of P. semisulcatus were identified in the Arabian Gulf waters in Iran (Jahromi et al., 2019). The proximity between the NW Red Sea samples of P. semisulcatus identified in the present study and some Strait of Hormuz samples, is indeed interesting. No studies were carried out to identify any relation between P. semisulcatus populations in the two locations; i.e. the Red Sea and the Arabian Gulf, at the prawn mitochondrial genetic level. However, a study pointed to genetic proximity between these two populations. Random amplified polymorphic DNA (RAPD), inter simple sequence repeat (ISSR), and simple sequence repeat (SSR) revealed a close relation between P. semisulcatus in Suez City in the NW Red Sea (Egypt) and that in Manifa area in the North Western part of the Arabian Gulf (Saudi Arabia) (Hassanien and Al‐Rashada 2019). A major drawback in the analyses of genetic connectivity among different world populations of P. semisulcatus is the absence of data from different areas in its range, including for example other locations in the Red Sea, the Gulf of Aden, and eastern Africa. Separate phylogeographic lineages and cryptic species are increasingly being reported within penaeid prawns all over the world, especially in the IWP and the related seas and gulfs. For example, the Kuruma prawn P. (Marsupenaeus) japonicus Spence Bate, 1888 was identified since 2014 to be a complex of two forms or species (Tsoi et al., 2014). One of them, P. japonicus or Form I, is solely located in the waters of East northern South China Sea areas. The other one, P. pulchricaudatus Stebbing, 1914 or Form II, is widely distributed in the South China Sea, western Indian Ocean, Australia, the Red Sea, and the Mediterranean (Tsoi et al., 2014). Also, three lineages were identified for the giant tiger prawn P. monodon Fabricius, 1798; one in the Western Pacific Ocean, other in the juncture of the Indian and Pacific Oceans, and the other is located in the Indian Ocean (Alam et al., 2016). The banana prawn P. merguiensis de Man, 1888 was suggested to include two related cryptic species (Hualkasin et al., 2003). The Pink shrimp F. subtilis (Pérez Farfante, 1967) was identified to contain two forms, that were previously named MI and MII (D'Incao et al., 1998; Maggioni et al., 2001), then identified as separate species under the names of F. subtilis and F. isabelae Tavares & Gusmão, 2016 (Tavares and Gusmão, 2016). Members of these species’ complexes showed differences that went deeper from only morphological variation, i.e. to a genetic transcriptional level. For instance, the P. pulchricaudatus exhibits higher background expression of heat shock proteins, which can be an indicator for superior thermal tolerance of P. pulchricaudatus over P. japonicus (Wang et al., 2020). Hence, it appears essential to provide more contribution for the taxonomy of different penaeids. This, in the era of major advances in molecular genetics, is crucial for proper species identification, physiological investigations, fisheries management, and species conservation.
5 Conclusions
In conclusion, P. semisulcatus sampled from three different locations of the NW Red Sea exhibited genetic variations among its populations. Such variations could be associated with the difference in environmental parameters of the different habits. In order to investigate how such genetic variations can affect the morphological and/or morphometric features of such populations, it is recommended to compare their morphometric features. Integrating these, further conservation and sustainable exploitation strategies can be applied for each population. Finally, the clear genetic disparity between the NW Red Sea population of P. semisulcatus and the other one at the opposite side of the Arabian tectonic plate exceeded the suggested threshold for cryptic species. Hence, future works to characterize the accurate range of this new lineage of P. semisulcatus are strongly recommended.
Author contributions
Conceptualization, Mohammed-Geba, K. & Yousif A, methodology, Mohammed-Geba K, software, Mohammed-Geba, K. & Yousif, A.; validation, Mohammed-Geba, K. & Yousif, A.; formal analysis, Mohammed-Geba, K. & Yousif, A.; investigation, Mohammed-Geba, K. & Yousif, A.; resources, Mohammed-Geba, K. & Yousif, A.; data curation, Mohammed-Geba, K.; writing—original draft preparation, Mohammed-Geba, K. & Yousif, A.; writing—review and editing, Mohammed-Geba, K. & Yousif, A.; visualization, Mohammed-Geba, K. & Yousif, A.; supervision, Mohammed-Geba, K. & Yousif, A.; project administration.
Data availability
The obtained COI sequences were deposited in the GenBank database under the accession numbers (acc. No. OM063021 - OM063096). All crude sequences are available upon requesting from the Editor or reviewers of Aquatic Living Resources. Also, the authors intend to deposit the sequences in the database recommended by the Editor-in-Chief of Aquatic Living Resources.
Conflicts of Interest
The authors declare no conflict of interest.
Supplementary Material
Supplementary Figure A: Main software and platforms applied for COI sequences´ data analyses in the present study: A: Chromas Lite 2.1, B: ClustalX2, C: IDS-BOLD, D: BLAST-GenBank, E: Dnasp6, F: PopArt 1.7, G: Arlequin 3.5.1.1., H: Mega11, I: MrBayes 3.2.1, J: BEAST, K:iToL. Access here
Acknowledgments
The authors would like to acknowledge the role Dr. M. Galal Khallaf (Cleveland Dental Institute, USA) for his thorough English Language review. We also would like to appreciate our deep thanks to the anonymous reviewers of and Editor of Aquatic Living Resources for their sincere guidance and comprehensive comments that contributed much to the manuscript.
References
- Alam MM. De Croos MDST, Pálsson S. 2017. Mitochondrial DNA variation reveals distinct lineages in Penaeus semisulcatus (Decapoda, Penaeidae) from the Indo‐West Pacific Ocean. Mar Ecol 38: e12406. [CrossRef] [Google Scholar]
- Alam MMM, Westfall KM, Pálsson S. 2016. Mitogenomic variation of Bangladesh Penaeus monodon (Decapoda, Penaeidae) and reassessment of its phylogeography in the Indo-West Pacific region. Hydrobiology 763: 249–265. [CrossRef] [Google Scholar]
- Bernardi G, Azzurro E., Golani D, Miller MR. 2016. Genomic signatures of rapid adaptive evolution in the bluespotted cornetfish, a Mediterranean Lessepsian invader. Mol Ecol 25: 3384–3396. [CrossRef] [PubMed] [Google Scholar]
- Boudouresque CF. 1999. The Red Sea-Mediterranean link: unwanted effects of canals, in Invasive species and biodiversity management, edited by O.T. Sandlund, P.J. Schei, A. Viken. Kluwer Academic Publishers, pp 213–228. [CrossRef] [Google Scholar]
- Darriba D, Taboada GL, Doallo R, Posada D. 2012. jModelTest 2: more models, new heuristics and parallel computing. Nat Methods 9: 772–772. [CrossRef] [PubMed] [Google Scholar]
- D'Incao, F., Delevedove G., Maggioni D.E., Maggioni R. 1998. Evidência genética da presença de Farfantepenaeus paulensis (Pérez Farfante, 1967) no litoral nordeste do Brasil (Decapoda: Penaeidae). Nauplius 6: 129–137. [Google Scholar]
- Ditty J.G. 2014. Sternal spines in penaeid postlarvae (Decapoda: Penaeidae): life-phase-specific and systematically significant? J Crust Biol 34: 618–628. [CrossRef] [Google Scholar]
- Dore I., Frimodt C. 1987. Penaeus semisulcatus, in An Illustrated Guide to Shrimp of the World 1987. Van Nostrand Reinhold, New York: Library of Congress Catalog Card Number 87-1399, pp. 168–169. [Google Scholar]
- Du X., Cai S., Yu C., Jiang X., Lin L., Gao T., Han Z. 2016. Population genetic structure of mantis shrimps Oratosquilla oratoria: testing the barrier effect of the Yangtze River outflow. Biochem Syst Ecol 66: 12–18. [CrossRef] [Google Scholar]
- El-Aiatt A.A., Shalloof K.A.S., Saber M.M. 2019. Bio-economic studies on the catch of Bardawil Lagoon, North Sinai, Egypt. Egypt J Aquat Res 45: 59–65. [CrossRef] [Google Scholar]
- El-Ganainy A.A., Yassien M.H. 2012. The population biology of penaeid prawns in the Gulf of Suez, Red Sea, Egypt. Mar Biol Res 8: 405–411. [CrossRef] [Google Scholar]
- Excoffier L., Lischer H.E. 2010. Arlequin suite ver 3.5: a new series of programs to perform population genetics analyses under Linux and Windows. Mol Ecol Res 10: 564–567. [CrossRef] [PubMed] [Google Scholar]
- Excoffier L., Smouse P.E., Quattro J.M. 1992. Analysis of molecular variance inferred from metric distances among DNA haplotypes: application to human mitochondrial DNA restriction data. Genetics 131: 479–491. [CrossRef] [PubMed] [Google Scholar]
- Folmer O., Black M., Hoeh W., Lutz R., Vrijenhoek R. 1994. DNA primers for amplification of mitochondrial cytochrome C oxidase subunit I from diverse metazoan invertebrates. Mol Mar Biol Biotechnol 3: 294–299. [PubMed] [Google Scholar]
- Fu Y.-X. 1997. Statistical tests of neutrality of mutations against population growth, hitchhiking and background selection. Genetics 147: 915–925. [CrossRef] [PubMed] [Google Scholar]
- Grant W.A.S., Bowen B.W. 1998. Shallow population histories in deep evolutionary lineages of marine fishes: insights from sardines and anchovies and lessons for conservation. J Hered 89: 415–426. [CrossRef] [Google Scholar]
- Halim S.A.A.A., Othman A.S., Akib N.A.M., Jamaludin N.A., Esa Y., Nor S.A.M. 2021. Mitochondrial Markers Identify a Genetic Boundary of the Green Tiger Prawn (Penaeus semisulcatus) in the Indo-Pacific Ocean. Zool Stud 60: e8. [PubMed] [Google Scholar]
- Han Z., Zhu W., Zheng W., Li P., Shui B. 2105. Significant genetic differentiation between the Yellow Sea and East China Sea populations of cocktail shrimp Trachypenaeus curvirostris revealed by the mitochondrial DNA COI gene. Biochem Syst Ecol 59: 78–84. [Google Scholar]
- Harpending H. 1994. Signature of ancient population growth in a low-resolution mitochondrial DNA mismatch distribution. Hum Biol 591–600. [PubMed] [Google Scholar]
- Hassanien H.A., Al-Rashada, Y. 2019. Estimation of genetic variation in green tiger prawn, Penaeus semisulcatus by using random amplified polymorphic DNA, inter simple sequence repeat and simple sequence repeat markers. Aquac Res 50: 1626–1633. [CrossRef] [Google Scholar]
- Hualkasin W., Sirimontaporn P., Chotigeat W., Querci J., Phongdara A. 2003. Molecular phylogenetic analysis of white prawns species and the existence of two clades in Penaeus merguiensis. J Exp Mar Biol Ecol 296: 1–11. [CrossRef] [Google Scholar]
- Jahromi S.T., Othman A.S., Rosazlina R. 2019. Morphometrics and mitochondrial DNA genes analysis suggest a new species of Penaeus (Crustacea: Penaeidae) from the Persian Gulf. Biochem Genet 57: 193–213. [CrossRef] [PubMed] [Google Scholar]
- Kasim N.S., Jaafar T.N.A.M., Piah R.M., Arshaad W.M., Nor S.A.M., Habib A., Abd Ghaffar M., Sung Y.Y., Danish-Daniel M., Tan M.P. 2020. Recent population expansion of longtail tuna Thunnus tonggol (Bleeker, 1851) inferred from the mitochondrial DNA markers. PeerJ 8: e9679. [CrossRef] [PubMed] [Google Scholar]
- Kiparissis S., Peristeraki P., Tampakakis K., Kosoglou I., Doudoumis V., Batargias C. 2018. Range expansion of a restricted lessepsian: westbound expansion breakthrough of Lagocephalus spadiceus (Richardson, 1844)(Actinopterygii: Tetraodontidae). Bioinvasions Rec 7. [Google Scholar]
- Lefebure T., Douady C.J., Gouy M., Gibert J. 2006. Relationship between morphological taxonomy and molecular divergence within Crustacea: Proposal of a molecular threshold to help species delimitation. Mol Phylogenet Evol 40: 435–447. [CrossRef] [PubMed] [Google Scholar]
- Maggioni R., Rogers A.D., MacLean N., D'Incao, F. 2001. Molecular phylogeny of western Atlantic Farfantepenaeus and Litopenaeus shrimp based on mitochondrial 16S partial sequences. Mol Phylogenet Evol 18: 66–73. [CrossRef] [PubMed] [Google Scholar]
- Mwakosya C.A., Mgaya Y.D., Jiddawi N.S. 2018. Genetic connectivity of Fenneropenaeus indicus (H. Milne Edwards 1837) among three prawn fishing grounds of Tanzanian coastal waters. Reg Stud Mar Sci 24: 107–112. [Google Scholar]
- Niamaimandi N., Aziz A., Siti Khalijah D., Che Roos S., Kiabi B. 2008. Reproductive biology of the green tiger prawn (Penaeus semisulcatus) in coastal waters of Bushehr, Persian Gulf. ICES Mar Sci 65: 1593–1599. [CrossRef] [Google Scholar]
- Pezy J.P., Baffreau A., Dauvin J.C. 2017. Records of two introduced Penaeidae (Crustacea: Decapoda) species from Le Havre Harbour, France, English Channel. Bioinvasions Rec 6: 363–367. [CrossRef] [Google Scholar]
- Rabaoui L., Lin Y.J., Maneja R.H., Qurban M.A., Abdurahiman P., Premlal P., Al-Abdulkader K., Roa-Ureta R.H. 2017. Nursery habitats and life history traits of the green tiger shrimp Penaeus semisulcatus (De Haan, 1844) in the Saudi waters of the Arabian Gulf. Fish Res 195: 1–11. [CrossRef] [Google Scholar]
- Ragonese S., Giusto G. 2011. Deep water occurrence of Penaeus semisulcatus De Haan, 1850, in the North Levant (Eastern Mediterranean Sea)(Crustacea: Decapoda: Penaeidae). Zool Middle East 54: 141–143. [CrossRef] [Google Scholar]
- Rambaut A., Drummond A.J., Xie D., Baele G., Suchard M.A. 2018. Posterior summarization in Bayesian phylogenetics using Tracer 1.7. System Biol 67: 901. [CrossRef] [PubMed] [Google Scholar]
- Ramos-Onsins S.E., Rozas J. 2002. Statistical properties of new neutrality tests against population growth. Mol Biol Evol 19: 2092–2100. [CrossRef] [PubMed] [Google Scholar]
- Ronquist F., Teslenko M., Van Der Mark P., Ayres D.L., Darling A., Höhna S., Larget B.L., Liang, Suchard M.A.; Huelsenbeck J.P. 2012. MrBayes 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space. Syst Biol 61: 539–542. [CrossRef] [PubMed] [Google Scholar]
- Rozas J., Ferrer-Mata A., Sánchez-DelBarrio J.C., Guirao-Rico S., Librado P., Ramos-Onsins S.E., Sánchez-Gracia A. 2017. DnaSP 6: DNA sequence polymorphism analysis of large data sets. Mol Biol Evol 34: 3299–3302. [CrossRef] [PubMed] [Google Scholar]
- Saher N., Matloob T., Aziz L., Awan A., Farooqi S. 2021. Adaptation of VB (Vartan Bhanji) in urban centers: tracking signs of cultural lag in an evolving Pakistan. Ilkogretim Online 20. [Google Scholar]
- Sharawy Z., Abbas E.M., Desouky M.G., Kato M. 2016. Descriptive analysis and molecular identification of the green tiger shrimp Penaeus semisulcatus (De Haan, 1844) in Suez Gulf, Red Sea. Int J Fish Aquat Stud 4: 426–432. [Google Scholar]
- Sharawy Z.Z., Abbas E.M., Khafage A.R., Galal-Khallaf A., Ismail R.F., Ahmed H.O., Mohammed-Geba K., Kato M. 2017. Descriptive analysis, DNA barcoding and condition index of Penaeids (Crustacea: Decapoda) from the Egyptian Mediterranean coast. Fish Res 188: 6–16. [CrossRef] [Google Scholar]
- Soyel H.I., Kumlu, M. 2003. The effects of salinity on postlarval growth and survival of Penaeus semisulcatus (Decapoda: Penaeidae). Turk J Zool 27: 221–225. [Google Scholar]
- Tajima F. 1989. Statistical method for testing the neutral mutation hypothesis by DNA polymorphism. Genetics 123: 585–595. [PubMed] [Google Scholar]
- Tamura K., Stecher G., Kumar S. 2021. MEGA11: molecular evolutionary genetics analysis version 11. Mol Biol Evol 38: 3022–3027. [CrossRef] [PubMed] [Google Scholar]
- Tavares C., Gusmão J. 2016. Description of a new Penaeidae (Decapoda: Dendrobranchiata) species, Farfantepenaeus isabelae sp. nov. Zootaxa 4171: 505–516. [CrossRef] [PubMed] [Google Scholar]
- Teodoro S.S.A., da Silva Cortinhas M.C., Proietti M.C., Costa R.C., Dumont L.F.C. 2020. High genetic connectivity among pink shrimp Farfantepenaeus paulensis (Pérez-Farfante, 1967) groups along the south-southeastern coast of Brazil. Estuar Coast Shelf Sci 232: 106488. [CrossRef] [Google Scholar]
- Tsoi K.H., Ma K.Y., Wu T.H., Fennessy S.T., Chu K.H., Chan T.Y. 2014. Verification of the cryptic species Penaeus pulchricaudatus in the commercially important kuruma shrimp P. japonicus (Decapoda: Penaeidae) using molecular taxonomy. Invertebr Syst 28: 476–490. [CrossRef] [Google Scholar]
- Walsh P.S., Metzger D.A., Higuchi R. 1991. Chelex 100 as a medium for simple extraction of DNA for PCR-based typing from forensic material. Biotechniques 10: 506–513. [PubMed] [Google Scholar]
- Wang P., Chen B., Zheng J., Cheng W., Zhang H., Wang J., Su Y., Xu P., Mao Y. 2020. Fine-Scale Population Genetic Structure and Parapatric Cryptic Species of Kuruma Shrimp (Marsupenaeus japonicus), Along the Northwestern Pacific Coast of China. Front Genet 11: 118. [CrossRef] [PubMed] [Google Scholar]
- Witt J.D.S., Threloff D.L., Hebert P.D.N. 2006. DNA barcoding reveals extraordinary cryptic diversity in an amphipod genus: Implications for desert spring conservation. Mol Ecol 15: 3073–3082. [CrossRef] [PubMed] [Google Scholar]
- WoRMS. 2021. Penaeus semisulcatus De Haan, 1844 [in De Haan, 1833-1850. Accessed at: http://marinespecies.org/aphia.php?p=taxdetails&id=107112 on 2021-05-28]. [Google Scholar]
- Xu D., Lou B., Shi H., Geng Z., Li S., Zhang Y. 2012. Genetic diversity and population structure of Nibea albiflora in the China Sea revealed by mitochondrial COI sequences. Biochem Syst Ecol 45: 158–165. [CrossRef] [Google Scholar]
- Yousif A. 2003. Diel variability of size and catch rate of three fish species and three penaeid prawns in the NW Red Sea trawl fishery. Fish Res 63: 265–274. [CrossRef] [Google Scholar]
Cite this article as: Mohammed-Geba K, Yousif A. 2022. COI gene sequencing reveals genetic population structuring and a separate lineage of the green tiger prawn Penaeus semisulcatus in the Gulf of Suez and the Bitter Lakes, Egypt. Aquat. Living Resour. 35: 4
All Tables
Accession numbers and locations for GenBank and BOLD-retrieved COI sequences of P. semisulcatus from different areas of the world.
Different haplotype diversity, nucleotide diversity, neutrality, and raggedness analyses values for Egyptian P. semisulcatus sampled from the Bitter Lakes and Gulf of Suez, Egypt.
Analysis of molecular variance (AMOVA) of Fst values for grouped P. semisulcatus COI sequences from different sampling locations in the NW of the Red Sea.
All Figures
 |
Fig. 1 The annual catch of ¨large shrimps¨ category, i.e. Penaeus semisulcatus and Penaeus japonicus in the Gulf of Suez, Egypt. Total numbers of catch in the period between 2014-2022 are shown in tons. Source: General Authority for fisheries resources development in Egypt (GAFRD), Suez branch (2022). |
| In the text | |
 |
Fig. 2 Sampling locations, shown as white points, for Penaeus semisulcatus from Bitter Lakes (N30º26'61.7'', E32 º 44'98.6'') and Gulf of Suez [Sadat (N29º45'74", E32º30'17) and Abo Regm (N28º43'35", E32º51'13)] at the North-Western Red Sea, Egypt. Photo from GoogleMapsTM (map credits are shown below the photo). |
| In the text | |
 |
Fig. 3 Mismatch distribution for COI-based pairwise distances of the North-Western Red Sea and Bitter lakes’ populations of P. semisulcatus. |
| In the text | |
 |
Fig. 4 Median-joining haplotype network for COI sequences in P. semisulcatus obtained from different areas in the North-West of the Red Sea. The branch length is proportional to the number of substitutions. Black transverse bars in the branches represent the polymorphic sites that differ among haplotypes. Circles represent haplotypes and their diameters are proportional to the haplotype frequencies. The Colour legend is shown on the down-right side of the graphic. |
| In the text | |
 |
Fig. 5 Median-joining haplotype network for COI sequences in P. semisulcatus from the Red Sea, the Arabian Gulf, and the Indo-West Pacific region (Bangladesh, Sri Lanka, Malaysia). The colour legend is shown on the down-right side of the graphic. |
| In the text | |
 |
Fig. 6 BI phylogenetic analysis for the samples collected in the present study. Posterior probabilities are shown above the branches. Green highlighted: Gulf of Suez P. semisulcatus haplotypes (Hap), and the closely related ones form the Arabian Gulf and the Mediterranean Sea. Red-highlighted: Indo-West Pacific (IWP) haplotypes. Tree outgroup: the Southern Horseshoe crab Tachypleus gigas. |
| In the text | |
 |
Fig. 7 Maximum-Likelihood (ML) phylogenetic tree for P. semisulcatus world haplotypes (Hap). Posterior probabilities are shown above the branches. Green highlighted: Gulf of Suez P. semisulcatus haplotypes (Hap), and the closely related ones form the Arabian Gulf and the Mediterranean Sea. Red-highlighted: Indo-West Pacific (IWP) haplotypes. Tree outgroup: the Southern Horseshoe crab Tachypleus gigas. |
| In the text | |
 |
Fig. 8 Distribution of genetic pairwise distances among Gulf of Suez, Arabian Gulf, and Indo-West Pacific P. semisulcatus COI haplotypes in comparison to those in the GenBank and BOLD databases. |
| In the text | |
Current usage metrics show cumulative count of Article Views (full-text article views including HTML views, PDF and ePub downloads, according to the available data) and Abstracts Views on Vision4Press platform.
Data correspond to usage on the plateform after 2015. The current usage metrics is available 48-96 hours after online publication and is updated daily on week days.
Initial download of the metrics may take a while.


