| Issue |
Aquat. Living Resour.
Volume 32, 2019
|
|
|---|---|---|
| Article Number | 16 | |
| Number of page(s) | 9 | |
| DOI | https://doi.org/10.1051/alr/2019017 | |
| Published online | 15 July 2019 | |
Research Article
Genetic diversity and population structure of small yellow croaker (Larimichthys polyactis) in the Yellow and East China seas based on microsatellites
1
Key Laboratory of Oceanic and Polar Fisheries, Ministry of Agriculture and Rural Affairs, East China Sea Fisheries Research Institute, Chinese Academy of Fishery Sciences, 200090 Shanghai, PR China
2
Department of Animal and Avian Sciences, University of Maryland, College Park, 20742 Maryland, USA
* Corresponding author: This email address is being protected from spambots. You need JavaScript enabled to view it.
, This email address is being protected from spambots. You need JavaScript enabled to view it.
Handling Editor: Costas S. Tsigenopoulos
Received:
25
October
2018
Accepted:
22
June
2019
Abstract
Small yellow croaker (Larimichthys polyactis), a member of family Sciaenidae, is mainly distributed in the northwestern Pacific Ocean. To assess the genetic diversity and population structure of this species across its range, we genotyped 150 L. polyactis individuals sampled in five locations along the coast of the Yellow and East China seas using 20 polymorphic microsatellites. A total of 499 alleles were detected at 20 loci across all individuals, and a relatively high level of genetic diversity was observed, with observed heterozygosity (Ho), expected heterozygosity (He) and polymorphic information content (PIC) ranging from 0.233 to 1.000, from 0.438 to 0.955, and from 0.367 to 0.953 per locus-location combination, respectively. Analysis of molecular variance (AMOVA) (FST = 0.00915, P < 0.001), pairwise FST, and corrected average pairwise differences indicated that there was extremely low, but statistically significant genetic differentiation among the studied populations. However, Bayesian assignment analysis revealed a high number of immigrants among populations and no obvious genetic differentiation. The Wilcoxon signed-rank test and mode-shift indicator of allele frequency distribution support the inferrence that L. polyactis had not experienced a recent genetic bottleneck. Overall, the results suggest that, despite low genetic differentiation in this species, the small yellow croaker forms a single panmictic population with high genetic variation and gene flow in the studied area. This study will provide useful information for conservation and sustainable exploitation of this important aquatic living resource.
Key words: Small yellow croaker / Larimichthys polyactis / Microsatellite / Genetic diversity / Population structure
© EDP Sciences 2019
1 Introduction
Small yellow croaker, Larimichthys polyactis, is an economically important demersal marine fish species belonging to family Sciaenidae in order Perciformes. It is native to the temperate waters of the northwestern Pacific Ocean, including the Bohai, Yellow, and East China seas (FishBase, 2012). It is an important marine food fish species in Asian countries, and it is the target of bottom trawling in China, Japan, and Korea (Jin et al., 2005). Due to overfishing and environmental degradation, its populations were reduced year after year in the 1970s, and the fisheries were on the verge of collapse by the early 1990s (Liu et al., 1990; Jin and Tang, 1996; Kim et al., 1997). A series of conservation programs have been implemented in China since the 1990s, including the establishment of non-fishing seasons and regions along the coast. Since then, the stock has recovered significantly and is continually increasing, with up to 438,837 t landed in 2010 (FAO, 2012). However, most of the fish landed (> 80%) were less than 1 year old, especially after 2000 (Jin et al., 2005; Yan et al., 2006). The stock of L. polyactis was therefore still quite fragile.
In addition, small yellow croaker is one of the most indispensable constituents in the ecosystem of northwest Pacific and has important ecological and evolutionary implication in native west Pacific fish. For example, on one hand, in the main distribution area of small yellow croaker, zooplankton is usually distributed with high density, but its abundance descended sharply in the autumn because of the strong feeding intensity of small yellow croaker and other plankton-feeding fishes in the summer (Li et al., 2007a). On the other hand, Yan et al. (2006) suggested that due to the general decline of fish stocks at the higher trophic level such as the small yellow croaker, their prey fish at lower trophic level have increased.
Due to the fact that some scientific experiments on the capture size of small yellow croaker are still under way and the opinions of different experts are controversial, till now, the management laws or rules on capture size of small yellow croaker have not been issued by the Chinese government. Under this situation, it is difficult to perform a strict capture size management for small yellow croaker and new management strategies are urgently needed for the sustainable growth and utilization of this species.
Since the 1990s, a series of conservation programs, including the establishment of non-fishing seasons and regions along the coast, has been implemented in China. Better knowledge about capture size and CPUE (Catch Per Unit Effort) is needed to improve present conservation programmes. Additionnally, information about genetic diversity and population structure can contribute to determine the Management Units (MUs) and the optimal strength of CPUE. These informations are very important basic knowledge to establish conservation programs.
For effective fishery management, we need a mass of biological and evolutionary data. Characterization of the genetic variation of this fish species is essential for conserving the genetic resource and reversing the population decline (Li et al., 2007b). Therefore, it is necessary to have a clear understanding of the intraspecific genetic diversity and population structure of the species.
Genetic diversity and population structure have vital importance in understanding and managing exploited populations (Palumbi, 2003; Muths et al., 2009). So far, some studies concerning the genetic diversity and population structure of L. polyactis have ever been reported. However, these studies yielded conflicting conclusions regarding the genetic diversity and population structure of this species. Studies using RAPD (random amplified polymorphic DNA) and AFLP (amplified fragment length polymorphism) markers (Meng et al., 2003; Han et al., 2009; Lin et al., 2009) revealed that there were at least three wild stocks. However, five mtDNA studies (Xiao et al., 2009; Wu et al., 2009; Peng et al., 2010; Kim et al., 2010; Wu et al., 2012) did not support the results of the two studies, showing no significant geographic structure in L. polyactis. These conflicting results have hampered management efforts for small yellow croaker. Thus, it is still necessary to investigate its genetic structure for further resource conservation and development of management strategies of this species.
Microsatellites or simple sequence repeats (SSR) are short (1-6 bp) repetitive DNA sequences, that are highly abundant and almost evenly distributed in genomes. They are generally neutral, highly polymorphic, co-dominant and easily scored with PCR (Goldstein and Schlotterer, 1999). Microsatellites have been extensively applied for genome mapping, population genetics, ecology, and evolution (Goldstein and Schlotterer, 1999; Herwerden et al., 1999). Over the past two decades, microsatellites have been regarded as one of the most useful tools for population genetics studies. In addition, microsatellites are reported to be more variable than RFLP or RAPD markers, and have been widely utilized in studies of fishery species (Liu and Cordes, 2004).
Population structure should be taken into account by regulatory authorities when considering sustainable exploitition of fish resource. The aim of this research was to examine the genetic diversity and population structure of the small yellow croaker in the Yellow Sea (YS) and East China Sea (ECS) to provide genetic information useful for the management of this important aquatic living resource. In the present study, we used twenty polymorphic microsatellite markers to examine the genetic diversity and population structure of five wild populations of L. polyactis along Yellow and East China Sea.
2 Materials and methods
All sampling sites were not privately-owned or protected, and field sampling did not involve protected species. The collection of the small yellow croaker complied with the guidelines of East China Sea Fisheries Research Institute.
2.1 Sampling and DNA extraction
A total of 150 wild L. polyactis individuals, i.e., 30 individuals of each population, were collected from five locations along the coast of the YS and ECS (Fig. 1). The five locations were Donggang (DG, 39°32′N, 124°5′E), Lianyungang (LYG, 34°22′N, 119°6′E), Shacheng (SC, 27°9′N, 120°24′E), Taizhou (TZ, 28°39′N, 121°48′E), and Yantai (YT, 37°18′N, 121°14′E), respectively. Muscle tissues were preserved in 100% ethanol and stored in the laboratory at room temperature until DNA extraction.
Total genomic DNA was extracted using traditional proteinase K and phenol-chloroform protocol. The quality of the extracted DNA was checked using 0.8% agarose gel electrophoresis, then the DNA was stored at −20°C for PCR amplification.
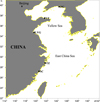 |
Fig. 1 Map of the Larimichthys polyactis sampled locations (indicated by circles). The abbreviations correspond to sampled location names, as indicated in text. DG: DongGang; LYG: LianYunGang; SC: ShaCheng; TZ: TaiZhou; YT: YanTai. |
2.2 Microsatellite genotyping
Twenty polymorphic microsatellite markers, i.e. XHY2, XHY4, XHY8, XHY9, XHY11, XHY12, XHY14, XHY15, XHY16, XHY17, XHY19, XHY21, XHY22, XHY23, XHY24, XHY25, XHY28, XHY30, XHY32, and XHY34 (Chen and Cheng 2013), were used for genotyping 150 L. polyactis individuals. Polymerase chain reaction (PCR) amplification was performed in a 15-µL reaction volume containing approximately 10–50 ng of genomic DNA, 0.6 μM of each primer, 7.5 μL 2 ×Taq PCR MasterMix (Tiangen), with the following thermal cycling conditions: an initial denaturation at 94 °C for 3 min, and then followed by 32 cycles of denaturing at 94 °C for 30 s, annealing at primer-specific annealing temperature (Ta , see Table 1 of Chen and Cheng, 2013) for 30 s, elongation at 72 °C for 30 s, and then followed by a final extension at 72 °C for 7 min.
The final PCR products were separated by electrophoresing in denaturing 6% polyacrylamide gels and visualized by silver staining. The allele sizes were estimated with a 10bp DNA ladder (Invitrogen) as reference.
AMOVA results of five Larimichthys polyactis populations.
2.3 Statistical analysis
2.3.1 Measurements of genetic diversity
The number of alleles (Na), number of effective alleles (Ne), observed heterozygosity (Ho), and expected heterozygosity (He) were obtained for each locus from each location using GenAlEx 6.0 (Peakall and Smouse, 2006). The diversity of each locus was evaluated with the polymorphic information content (PIC) using CERVUS version 3.0.3 (Marshall et al., 1998). GENEPOP on the Web (http://genepop.curtin.edu.au/, Raymond and Rousset, 1995) was used to assess the deviations from Hardy-Weinberg equilibrium (HWE) and linkage disequilibrium (LD). HWE and LD exact tests were tested using the Markov Chain method (10,000 dememorization steps, 100 batches, 5000 iterations). The inbreeding coefficient (FIS) was evaluated for each population at each locus using the software Arlequin version 3.1 (Excoffier et al., 2005).
2.3.2 Estimation of population differentiation
To study population structure, population-level pairwise FST was analyzed using a permutation procedure with 10,000 replicates. Corrected average pairwise differences (PiXY−(PiX+PiY)/2) and analysis of molecular variance (AMOVA) were calculated using Arlequin 3.1 (Excoffier et al., 2005). With the AMOVA, the L. polyactis populations were treated as a single group, rather than several hierarchically nested groups, because there was no obvious ocean barrier. In AMOVA, total variance is partitioned into two separate components, which describe the proportions of the total variance within and among populations. Sequential Bonferroni correction was used to adjust the significance levels of multiple tests (Rice, 1989).
To examine the genetic relationships among populations, the matrix of Nei's genetic distances among pairwise locations (Nei, 1978) was calculated using GenAlEx 6.0 (Peakall and Smouse, 2006). Then, an unweighted pair-group mean analysis (UPGMA) tree was constructed based on Nei's genetic distance matrices using MEGA version 5.0 software (Tamura et al., 2011). In order to provide a visual representation of population subdivision, principal components analysis (PCA) was performed in GenAlEx 6.0 (Peakall and Smouse, 2006).
2.3.3 Migration and demographical bottleneck
To evaluate individual migrant events across the marine current and investigate population mixing, an assignment test was implemented by GENECLASS 2.0 (Piry et al., 2004) according the probability of each individual assigned to a particular population based on multilocus genotype data. Individuals were assigned to the inferred genetic cluster to which they had the highest membership value. Migration events were identified as fishes assigning to inferred genetic clusters that were different from the location where they were sampled. First-generation (F0) migrant detection was also calculated using the ratio of the likelihood value from the population where the individual was sampled, divided by the likelihood value among all population samples, including the population where the individual was sampled from (L_home/L_max) as the test statistic. The assignment tests were computed using Bayesian approach (Rannala and Mountain, 1997) under a 0.05-threshold and employing the Monte Carlo resampling method with 10,000 simulated individuals and the type I error rate of 0.1 (Paetkau et al., 2004).
In addition, we detect recent population declines by computing the heterozygosity excess statistic using the computer program BOTTLENECK (Cornuet and Luikart, 1996) with two methods. The first method basing on the principle of heterozygosity excess was executed under two different mutation models: the two-phase mutation model (TPM) and stepwise mutation model (SMM), where 95 and 90% single-step mutations, and 5 and 10% multiple steps mutations with 1000 simulation iterations were set as recommended by Piry et al. (1999). The SMM was used because microsatellite loci appear to evolve under a mutation model that is more similar to the SMM than the infinite allele model (IAM) (Valdes et al., 1993; Shriver et al., 1993). The second method, the mode-shift test (Luikart and Cornuet, 1998), was used to detect a potentially indicator of bottlenecked population using an L-shaped distribution of allele frequency as mutation-drift equilibrium.
3 Results
3.1 Genetic diversity
All the 20 microsatellite markers were well amplified in the five populations of small yellow croaker. No significant linkage between any pair of loci after sequential Bonferroni corrections (minimum adjusted alpha = 0.000053), which indicates that the assortment of alleles was independent at the 20 loci. Hence, all the markers were considered to be independently segregating for further analyses. Of the 100 HWE tests across all 20 loci, 88 were conformed to HWE within each sample location after applying sequential Bonferroni corrections (minimum adjusted alpha = 0.00050) (Rice, 1989). When each location was analyzed separately, there was no evidence for stuttering and allelic dropout in any of the loci assessed by Micro-Checker 2.2.3 (Van Oosterhout et al., 2004).
All 20 loci were polymorphic in all five populations of L. polyactis, with high allelic diversity and heterozygosity (Table S1). A total of 499 alleles were detected at the 20 microsatellite loci across the five populations. XHY2 was the most polymorphic microsatellite with 50 alleles, while XHY30 was the least variable one with 7 alleles. Among the five populations, a total of 111 private alleles were detected. Except for SC, with only 12 private alleles detected among the 20 markers, the numbers of private alleles in the other populations were all above 20. The allele frequencies for the 20-microsatellite loci did not vary significantly among the different populations (see examples XHY8 and XHY25 in Fig. S1).
For the five populations, the average polymorphism information content (PIC) ranged from 0.367 to 0.953 per locus-location combination, and from 0.831 (DG) to 0.845 (TZ) per location. Observed and expected heterozygosities (Ho and He) ranged from 0.233 to 1.000 and from 0.438 to 0.955 per locus-location combination, while from 0.762 (YT) to 0.799 (DG) and from 0.845 (DG) to 0.858 (TZ) per location, respectively (Table S1). Within sampling locations, the mean expected heterozygosities (He) were consistently higher than the observed one (Ho) across all loci, which revealed a deficit of heterozygosity among the samples. The inbreeding coefficient (FIS) ranged from −0.156 to 0.480 per locus-location combination and from 0.071 (DG) to 0.126 (YT) per location, with an average of 0.097 as a whole. All the indexes suggest that there is low inbreeding in these five L. polyactis populations.
3.2 Population structure
Significant genetic heterozygosity among the five populations was indicated by results of AMOVA analysis showed that 0.91% of total genetic variation came from among population variation, while within population variation explained 99.09% of total variation. Genetic differentiation between populations was analyzed using FST and corrected average pairwise differences. Although the overall FST value over all locations and loci was statistically different from zero (Fixation Index FST = 0.00915, P < 0.001), the genetic differentiation was still low, because the FST values were much lower than 0.05 (Tables 1 and 2). Pairwise FST comparing population pairs ranged from 0.004 (TZ and YT) to 0.015 (DG and LYG) (Table 2). All estimates of pairwise FST, except the comparison between TZ and YT, were significantly different from zero (P < 0.05). The highest genetic differentiation between populations, using both FST and corrected average pairwise difference, was between the populations DG and LYG, while the lowest differentiation was between TZ and YT (Table 2). The results of FST and corrected average pairwise difference indicated that there was extremely low but statistically significant genetic differentiation among the populations.
UPGMA phylogenetic trees were constructed on the basis of Nei's genetic distance matrix. Five groups were gathered into three main clusters: the DG population formed Cluster I; Cluster II included population SC; and the remaining populations formed Cluster III (Fig. S2). Furthermore, the PCA analysis also showed the DG population to be genetically distinct from the other four locations (Fig. 2). These results were consistent with the corrected average pairwise differences and FST tests.
Estimates of pairwise FST values (below diagonal) and corrected average pairwise differences (PiXY−(PiX+PiY)/2 (above diagonal) among five Larimichthys polyactis populations detected by 20 microsatellites.
 |
Fig. 2 Principal component analysis (PCA) of the five populations of Larimichthys polyactis. |
3.3 Migration
The inference of recent migration calculated in GENECLASS revealed a high number of immigrants among populations and ten of them were present generation (F0) immigrants (Table 3). The LYG, SC, TZ, and YT populations showed higher proportions of assignment probability, with the YT population showing the highest assignment probability with 24 immigrants assigned to other populations, while the DG population exhibited a lower number with 8 immigrants. The GENECLASS results also revealed that 36 individuals were identified as migrants from the ECS (SC, TZ) to the YS (LYG, DG, TY), whereas 46 from the YS were assigned to the ECS (Table 3). The results indicated that almost all the populations and individuals were admixed, showing that generally limited divergence throughout the examined range of L. polyactis.
Number of immigrants assigned to a given population performed with Geneclass.
3.4 Demographic bottleneck
Analysis of recent population declines of L. polyactis was detected using the Wilcoxon signed-rank test in BOTTLENECK under two models of microsatellite evolution (SMM and TPM) with two sets of parameters (Table S2). The probability values of the bottleneck test ranged from 0.24900 to 0.99928, all above 0.05, suggesting no significant excess of heterozygosity. It indicated that there was no genetic bottleneck in the five populations due to mutation-drift equilibrium. In addition, the mode-shift test showed that all populations were in normal L-shaped pattern of the allele frequency distributions (Table S2), revealing the lack of population declines in the recent history of L. polyactis. Therefore, we infer that none of the studied populations have experienced a recent genetic bottleneck.
4 Discussion
In this study, we examined the genetic diversity and population structure of five wild populations of L. polyactis using 20 highly polymorphic microsatellites. We observed low but significant genetic differences in the spatial distribution of small yellow croaker in the YS and the ECS. The results suggested a high level of population genetic diversity of L. polyactis (Na:15.05-16.70, PIC:0.831-0.845, Ho:0.762-0.799, and He: 0.845-0.858), which is in line with the previous studies that showed a high level of genetic diversity in L. polyactis using RAPD, ISSR, and mtDNA markers (Meng et al., 2003; Xu et al., 2005; Wu et al., 2009; Peng et al., 2010, Wu et al., 2012; Zheng et al., 2012), and very close to the result of Kim et al. (2012) with He ranged from 0.776 to 0.947. This evaluation of the genetic diversity and population structure of L. polyactis is both necessary and significant. Across the five sample locations, the average number of alleles was 24.95 per locus, higher than those (11.3/locus, 15/locus, 10.5/locus) found in three other marine fishes (DeWoody and Avise, 2000; Li et al., 2006, Zeng et al., 2012), suggesting that high genetic variation is available in L. polyactis. In comparison to other marine fish species (DeWoody and Avise, 2000; Li et al., 2006; Zeng et al., 2012), we also found the L. polyactis exhibited higher expected heterozygosity. Overall, the gene diversity in the studied populations of L. polyactis was above average among marine fishes. The levels of genetic diversity of L. polyactis from the YS and ECS were almost equal (Table S1), showing no obvious geographic predominance. This phenomenon may be caused by its wide distribution and diverse inhabitation environments. All these data indicated that the fish from the five wild populations in the YS and ECS contained high allelic and gene diversity. Future studies on genetic diversity of small yellow croaker using microsatellites should include samples from Bohai Sea to get a more comprehensive picture of genetic variation of this species.
The amount of effective alleles (Ne) was obviously less than that of the observed alleles (Na) (Table S1), indicating that some alleles were lost in the populations studied in this study. Furthermore, many private alleles were found at a very low frequency in this study. The alleles with low frequency may easily go extinct (Brookes et al., 1997). However, the number of alleles is very important to maintain populations because it reflects the necessary spectrum of genotypes for adaptive response to changing environments (Thai et al., 2007). The loss of genetic diversity, or even that of some private alleles, may prevent future improvements via selection of the species to a certain extent. The most likely explanation for the loss of alleles may be that the wild population was under stress from overfishing and deterioration in the environment resulting in less recruitment (Alam and Islam, 2005), and the limited sampling of populations may also play a role here. The level of the genetic diversity is high, we should still pay attention to the protection of this species. In order to conserve the genetic resource, we could strengthen the conservation and supervision of wild resources and forbid overfishing in the natural waters.
Generally, marine species are capable of migration over long distances and show little or no genetic differentiation in geographic scales because of long distance pelagic dispersal potential during planktonic egg, larval, or adult history stages coupled with an absence of physical barriers to movement between ocean basins or adjacent continental margins (Hewitt, 2000). The small but highly significant values of pairwise FST in the present study indicated that the small yellow croaker has only subtle genetic differentiation among populations, with more than 99.09% of the total genetic variation among individuals within populations (Table 1). UPGMA and PCA analysis indicated that the population DG more genetically different from other populations. In all, these results revealed no significant genetic structure or population subdivision throughout the examined range of sampled locations. Low FST value suggested relatively high gene flow among different locations, which means that the small yellow croaker was likely a single panmictic population.
Based on morphology and ecological approaches, scientists identified three wild stocks, North Yellow Sea, South Yellow Sea, and East China Sea populations, existed across the distribution of L. polyactis. Furthermore, the earlier studies using RAPD (Meng et al., 2003) and AFLP markers (Han et al., 2009; Lin et al., 2009) did support the idea of three wild stocks. However, our study results, as well as the five mtDNA studies in the similar areas (Xiao et al., 2009; Wu et al., 2009; Peng et al., 2010; Kim et al., 2010; Wu et al., 2012) were inconsistent with those of the studies which showed significant geographic structure in L. polyactis. The reasons lead to this divergence may include intrinsic variability of the population, sample size, spatial replication, and the number and characteristics of the marker loci used (Curley and Gillings, 2009).
Ikeda (1964) found that recruitment phenomenon existed among wild stocks of L. polyactis based on the investigation of fishery status. The migration analysis, calculated in GENECLASS, found a high number of potential migrants within the YS and ECS groups. A complex system of surface currents is a more likely influencing factor that impeded the connectivity of gene flow. The China Coastal Current and Yellow Sea Warm Current, including outflow of water from the Yellow Sea to the East China Sea along the China coast and inflow from the East China Sea to the Yellow Sea along the west coast of Korea (Li et al., 2000), respectively, could contribute to mixing of the ECS and YS groups. Additionally, small yellow croaker migrate respectively towards the coastal waters of Jiangsu (the northern waters of Yangtze estuary) and Zhejiang Provinces (north Zhejiang coast and Yangtze estuary waters) to spawn. After spawning, the spent fish remain in these areas to feed, whereas the juveniles dispersed to the nursery grounds in both the YS and the ECS until autumn (Lin, 1964; Wang et al., 1965; Zhao et al., 1987; Liu, 1990).
The past demographic history associated with the distribution of the species due to historical vicariance influence the detection of the recent bottleneck events. Bottleneck analysis revealed that the five populations have not undergone a recent bottleneck. The result indirectly supported the high degree of gene flow and relatively low genetic differentiation in L. polyactis.
Dispersal of breeding individuals between populations may be the main determinant of the genetic structure of population beside geographical barriers (Coulon et al., 2006). Because small yellow croaker is a migratory fish, its ability to spread is relatively strong, which is conducive to gene flow. In this study, genetic diversity of small yellow croaker was relatively high and we could infer that gene flow to and from the study sites was possible. Relatively high levels of heterozygosity found in this study suggest that either the area and population are large enough to sustain genetic diversity, or the population is still connected to other subpopulations (Morandin et al., 2014).
In conclusion, our study demonstrated that the five wild populations of small yellow croaker (L. polyactis) along the YS and ECS contained high levels of genetic diversity but exhibited quite low differentiation. The data generated in this study provided useful information for conservation and exploitation of this fishery resource. The Management unit (MU) concept, proposed by Moritz (1994), was defined as a conservation unit that had statistically significant divergence in allele frequencies (nuclear or mitochondrial). Since no obvious genetic differentiation and genetic structure between different geographical populations of small yellow croaker, we suggest that the populations of small yellow croaker in the Yellow Sea and the East China Sea should be deemed as one Management Unit (MU) with the same management strategy. Therefore, the small yellow croaker in the YS and ECS could be considered as one unit, which provides a guideline for further effective conservation and management of this important aquatic living resource.
Supplementary Material
Fig. S1 Allele frequencies at two selected microsatellite loci (XHY8 and XHY25) for the five populations: DG, LYG, SC, TZ, YT. The X axis shows allele sizes, while frequencies are indicated on the Y axis.
Fig. S2 Unweighted Pair-group Method with Arithmetic Means tree (UPGMA) among 5 populations of Larimichthys polyactis constructed with the Nei’s genetic distances.
Table S1 Statistics for genetic variation at 20 microsatellite loci in five populations of Larimichthys polyactis collected from the Yellow Sea and East China Sea.
Table S2 P-values of bottleneck tests for detecting the recent population declines of Larimichthys polyactis using two phased mutation model (TPM), stepwise mutation model (SMM) and mode shift indicator.
Access Supplementary MaterialAcknowledgments
We would like to thank three anonymous reviewers, one anonymous Associate Editor, and Dr. Pierre Boudry, the Editor-in-Chief of Aquatic Living Resources for their valuable comments on this paper. We would also like to thank Mr Hao Huang, Ms Yuxia Zhu, and Ms Yushuang Ge for assistance in sample and data collection. This research was supported by Central Public-interest Scientific Institution Basal Research Fund, ECSFR, CAFS (No.2012M09) and Science and Technology Commission of Shanghai Municipality (Shanghai Natural Science Foundation No. 13ZR1449800). All the sampling sites were not privately-owned or protected, and field sampling did not involve protected species. The collection of the small yellow croaker fish complied with the guidelines of East China Sea Fisheries Research Institute.
References
- Alam MS, Islam MS. 2005. Population genetic structure of Catla catla(Hamilton) revealed by microsatellite DNA markers. Aquaculture 246: 151–160. [Google Scholar]
- Brookes MI, Graneau YA, King P, Rose OC, Thomas CD, Mallet JLB. 1997. Genetic analysis of founder bottlenecks in the rare British butterfly Plebejus argus . Conserv Biol 11: 648–661. [Google Scholar]
- Chen WM, Cheng QQ. 2013. Development of thirty-five novel polymorphic microsatellite markers in Larimichthys polyactis (Perciformes: Sciaenidae) and cross-species amplification in closely related species, Pseudosciaena crocea . Biochem Syst Ecol 47: 111–115. [Google Scholar]
- Cornuet JM, Luikart G. 1996. Description and power analysis of two tests for detecting recent population bottlenecks from allele frequency data. Genetics 144: 2001–2014. [PubMed] [Google Scholar]
- Coulon A, Guillot G, Cosson J. 2006. Genetic structure is influenced by lansdcape features. Empirical evidence from a roe deer population. Mol Ecol 15: 1669–1679. [CrossRef] [PubMed] [Google Scholar]
- Curley BG, Gillings MR. 2009. Population connectivity in the temperate damselfish Parma microlepis: analysis of genetic structure across multiple spatial scales. Mar Biol 156(3): 381–393. [Google Scholar]
- DeWoody JA, Avise JC. 2000. Microsatellite variation in marine, freshwater and anadromous fishes compared with other animals. J Fish Biol 56: 461–473. [Google Scholar]
- Excoffier L, Laval G, Schneider S. 2005. Arlequin version 3.0: an integrated software package for population genetics data analysis. Evol Bioinform 1: 47–50. [CrossRef] [Google Scholar]
- FAO. 2012. http://www.fao.org/fishery/species/2362/en. Accessed 10 Oct 2012. [Google Scholar]
- FishBase. 2012. http://www.fishbase.org/home.htm. Accessed 20 Dec 2012. [Google Scholar]
- Goldstein DB, Schlotterer C. 1999. Microsatellites: evolution and applications. Oxford University Press, Oxford. [Google Scholar]
- Han ZQ, Lin LS, Shui BN, Gao TX. 2009. Genetic diversity of small yellow croaker Larimichthys polyactis revealed by AFLP markers. Afr J Agr Res 4(7): 605–610. [Google Scholar]
- Herwerden L, Blair D, Agatsuma T. 1999. Genetic diversity in parthenogenetic triploid Paragonimus westermani . Int J Parasi 29(9): 1477–1482. [CrossRef] [Google Scholar]
- Hewitt GM. 2000. The genetic legacy of the Quaternary ice ages. Nature 405: 907–913. [CrossRef] [PubMed] [Google Scholar]
- Ikeda I. 1964. Studies on the fisheries biology of the Yellow Croaker in the East China and the Yellow Seas. Seikai Reg Fish Res Lab 31: 1–81. [Google Scholar]
- Jin X, Zhao X, Meng T, Cui Y. 2005. Biology resources and habitat environment in Yellow and Bohai Seas. Science Press, Beijing, (in Chinese). [Google Scholar]
- Jin X, Tang Q. 1996. Changes in fish species diversity and dominant species composition in the Yellow Sea. Fish Res 26: 337–352. [Google Scholar]
- Kim JK, Min GS, Yoon M, Kim Y, Choi JH, Oh TY, Ni Y. 2012. Genetic structure of Larimichthys polyactis (Pisces: Sciaenidae) in the Yellow and East China Seas inferred from microsatellite and mitochondrial DNA analyses. Animal Cells Syst 16(4): 313–320. [CrossRef] [Google Scholar]
- Kim S, Jung S, Zhang CI. 1997. The effect of seasonal anomalies of seawater temperature and salinity on the fluctuation in yields of small yellow croaker, Larimichthys polyactis, in the Yellow Sea. Fish Oceanogr 6: 1–9. [Google Scholar]
- Kim JK, Kim YH, Kim MJ, Park JY. 2010. Genetic diversity, relationships and demographic history of the small yellow croaker, Larimichthys polyactis (Pisces: Sciaenidae) from Korea and China inferred mitochondrial control region sequence data. Anim Cells Syst 14(1): 45–51. [CrossRef] [Google Scholar]
- Li DY, Kang DH, Yin QQ, Sun XW, Liang LQ. 2007b. Microsatellite DNA marker analysis of genetic diversity in wild common carp (Cyprinus carpio L.) populations. J Genet Genom 34(11): 984–993. [CrossRef] [Google Scholar]
- Li J, Feng F, Yue GH. 2006. Twelve novel polymorphic microsatellites in a marine fish species, yellow croaker Larimichthys polyactis . Mol Ecol Notes 6: 188–190. [Google Scholar]
- Li JS, Yan LP, Li HY, Yang Li. 2007a. On the relationship between quantity distribution of small yellow croaker (Larimichthys polyactis Bleeker) and zooplankton in Southern Yellow Sea and the Northern East China Sea in summer and autumn. Marine Fisheries 29(1): 31–37. [Google Scholar]
- Li NS, Zhao SL, Wasiliev B. 2000. Geology of marginal sea in the Northwest Pacific. Heilongjiang Education Press, Harbin, (in Chinese). [Google Scholar]
- Lin XZ. 1964. The study on biological measurement of population of small yellow croaker. In: The Proceedings of Marine Fisheries Resources. Agriculture Press, Beijing, (in Chinese). [Google Scholar]
- Lin LS, Ying YP, Han ZQ, Xiao YS, Gao TX. 2009. AFLP analysis on genetic diversity and population structure of small yellow croaker Larimichthys polyactis . Afr J Biotechnol 8(12): 2700–2706. [Google Scholar]
- Liu X, Wu J, Han G. 1990. Investigation and division of the Yellow Sea and Bohai Sea Fishery Resources. Ocean Press, Beijing, (in Chinese). [Google Scholar]
- Liu XS. 1990. The Fisheries Resources Investigation and Division of the Yellow and Bohai Seas. Ocean Press, Beijing, (in Chinese). [Google Scholar]
- Liu ZJ, Cordes JF. 2004. DNA marker technologies and their applications in aquaculture genetics. Aquaculture 238: 1–137. [Google Scholar]
- Luikart G, Cornuet JM. 1998. Empirical evaluation of a test for identifying recently bottlenecked populations from allele frequency data. Conserv Biol 12: 228–237. [Google Scholar]
- Marshall TC, Slate J, Kruuk LEB, Pemberton JM. 1998. Statistical confidence for likelihood-based paternity inference in natural populations. Mol Ecol 7: 639–655. [CrossRef] [PubMed] [Google Scholar]
- Meng ZN, Zhuang ZM, Jin XS, Tang QS, Su YQ. 2003. Genetic diversity in small yellow croaker (Larimichthys polyactis) by RAPD analysis. Biodivers Sci 11(3): 197–203. [Google Scholar]
- Morandin C, Loveridge AJ, Segelbacher G. 2014. Gene flow and immigration: genetic diversity and population structure of lions (Panthera leo) in Hwange National Park, Zimbabwe. Conserv Genetics 15(3): 697–706. [CrossRef] [Google Scholar]
- Moritz C. 1994. Defining “evolutionary significant units” for conservation. Trends Ecol Evol 9: 373–375. [CrossRef] [PubMed] [Google Scholar]
- Muths D, Grewe P, Jean C. 2009. Genetic population structure of the Swordfish (Xiphias gladius) in the southwest Indian Ocean: sex-biased differentiation, congruency between markers and its incidence in a way of stock assessment. Fish Res 97: 263–269. [Google Scholar]
- Nei M. 1978. Estimation of average heterozygosity and genetic distance from a small number of individuals. Genetics 89: 583–590. [PubMed] [Google Scholar]
- Paetkau D, Slade R, Burden M, Estoup A. 2004. Direct, real-time estimation of migration rate using assignment methods: a simulation-based exploration of accuracy and power. Mol Ecol 13: 55–65. [CrossRef] [PubMed] [Google Scholar]
- Palumbi SR. 2003. Population genetics, demographic connectivity, and the design of marine reserves. Ecol Appl 13: S146–S158. [Google Scholar]
- Peakall R, Smouse PE. 2006. GENALEX 6: genetic analysis in Excel. Population genetic software for teaching and research. Mol Ecol Notes 6: 288–295. [Google Scholar]
- Peng B, Zhang Q, Zhao S, Le XL. 2010. Genetic diversity analysis of Larimichthys polyactis in coastal waters of China based on cytochrome B gene. Guangdong Agr Sci (China) 2: 131 –135. [Google Scholar]
- Piry S, Alapetite A, Cornuet JM, Paetkau D, Baudouin L, Estoup A. 2004. GeneClass2: a software for genetic assignment and first-generation migrant detection. J Hered 95: 536–539. [CrossRef] [PubMed] [Google Scholar]
- Piry S, Luikart G, Cornuet JM. 1999. BOTTLENECK: a computer program for detecting recent reductions in the effective population size using allele frequency data. J Hered 90: 502–503. [Google Scholar]
- Rannala B, Mountain JL. 1997. Detecting immigration by using multilocus genotypes. Proc Natl Acad Sci 94: 9197–9201. [Google Scholar]
- Raymond M, Rousset F. 1995. Genepop (version-1.2) population genetics software for exact tests and ecumenicism. J Hered 86: 248–249. [Google Scholar]
- Rice WR. 1989. Analyzing tables of statistical tests. Evolution 43: 223–225. [CrossRef] [PubMed] [Google Scholar]
- Shriver MD, Jin L, Chakraborty R, Boerwinkle I. 1993. VNTR allele frequency distributions under the stepwise mutation model − a computer simulation approach. Genetics 134: 983 –993. [PubMed] [Google Scholar]
- Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S. 2011. MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28(10): 2731 –2739. [CrossRef] [PubMed] [Google Scholar]
- Thai BT, Burridge CP, Austin CM. 2007. Genetic diversity of common carp (Cyprinus carpio) in Vietnam using four microsatellite loci. Aquaculture 269: 74–186. [Google Scholar]
- Valdes AM, Slatkin M, Freimer NB. 1993. Allele frequencies at microsatellite loci − the stepwise mutation model revisited. Genetics 133: 737–749. [PubMed] [Google Scholar]
- Van Oosterhout C, Hutchinson WF, Wills DPM, Shipley P. 2004. MICRO-CHECKER: software for identifying and correcting genotyping errors in mirosatellite data. Mol Ecol Notes 4: 535–538. [Google Scholar]
- Wang YG, Ma ZY, You HB. 1965. The preliminary study on migration and distribution of small yellow croaker. In: The Proceedings of Marine Fisheries Resources. Agriculture Press, Beijing, (in Chinese). [Google Scholar]
- Wu R, Liu S, Zhuang Z, Jin XS, Su YQ, Tang QS. 2009. Population genetic structure of small yellow croaker based on cytochrome b sequences. Prog Nat Sci (China) 19: 924–930. [Google Scholar]
- Wu R, Liu S, Zhuang Z. 2012. Population genetic structure and demographic history of small yellow croaker, Larimichthys polyactis (Bleeker, 1877), from coastal waters of China. Afr J Biotech 11: 12500–12509. [Google Scholar]
- Xiao YS, Zhang Y, Gao TX, Yanagimoto T, Yabe M, Sakurai Y. 2009. Genetic diversity in the mtDNA control region and population structure in the small yellow croaker, Larimichthys polyactis . Environ Biol Fish 85: 303–314. [CrossRef] [Google Scholar]
- Xu GP, Zhong XM, Ding YP, Liu PT, Tang JH, Xu P. 2005. The research on genetic diversity of Larimichthys polyactis population from the southern part of the Yellow Sea. Mar Sci (China) 29: 34–38. [Google Scholar]
- Yan L, Hu F, Ling J, Li S. 2006. Study on age and growth of Larimichthys polyactis in the East China Sea. Ocean Univ China 36: 95–100. [Google Scholar]
- Yan LP, Li JS, Shen DG, Yu LF, Ling LY. 2006. Variations in diet composition and feeding intensity of small yellow croaker Larimichthys polyactis Bleeker in the southern Yellow Sea and northern East China Sea. Marine Fisheries 28(2): 162–170. [Google Scholar]
- Zeng LY, Cheng QQ, Chen XY. 2012. Microsatellite analysis reveals the population structure and migration patterns of Scomber japonicus (Scombridae) with continuous distribution in the East and South China Seas. Biochem Syst Ecol 42: 83–93. [Google Scholar]
- Zhao CY, Chen YF, Hong GC. 1987. Fishery resources investigation in East China Sea. East China Normal University Publisher, Shanghai, (in Chinese). [Google Scholar]
- Zheng WJ, Lai YH, You QY, Qin QH, Zhu SH. 2012. Genetic diversity of Larimichthys polyactis in Zhoushan based on mitochondrial DNA D-loop region sequences. Zool Res 33: 329–336. [CrossRef] [PubMed] [Google Scholar]
Cite this article as: Cheng Q, Chen W, Ma L. 2019. Genetic diversity and population structure of small yellow croaker (Larimichthys polyactis) in the Yellow and East China seas based on microsatellites. Aquat. Living Resour. 32: 16
All Tables
Estimates of pairwise FST values (below diagonal) and corrected average pairwise differences (PiXY−(PiX+PiY)/2 (above diagonal) among five Larimichthys polyactis populations detected by 20 microsatellites.
All Figures
 |
Fig. 1 Map of the Larimichthys polyactis sampled locations (indicated by circles). The abbreviations correspond to sampled location names, as indicated in text. DG: DongGang; LYG: LianYunGang; SC: ShaCheng; TZ: TaiZhou; YT: YanTai. |
| In the text | |
 |
Fig. 2 Principal component analysis (PCA) of the five populations of Larimichthys polyactis. |
| In the text | |
Current usage metrics show cumulative count of Article Views (full-text article views including HTML views, PDF and ePub downloads, according to the available data) and Abstracts Views on Vision4Press platform.
Data correspond to usage on the plateform after 2015. The current usage metrics is available 48-96 hours after online publication and is updated daily on week days.
Initial download of the metrics may take a while.


